TF_ChIP-seq_ENCSR000ARE
| Id: | TF_ChIP-seq/ENCSR000ARE |
| Type: | position_score |
| Version: | 0 |
| Summary: |
TFChIP-seq ENCSR000ARE [biosamplesummary="Homo sapiens mammary epithelial cell female adult (50 years)" and target="EZH2"] |
| Description: |
status: released biological_replicates: Rep 1, Rep 2 summary: female adult (50 years) output_type: optimal IDR thresholded peaks audit_internal_action: Archived analysis {ENCAN966EMB|/analyses/ENCAN966EMB/} has in progress subobject quality standard {encode3-tf-chip|/quality-standards/encode3-tf-chip/} audit_warning: Replicate concordance in ChIP-seq experiments is measured by calculating IDR values (Irreproducible Discovery Rate). ENCODE processed IDR thresholded peaks files {ENCFF177LBB|/files/ENCFF177LBB/}, {ENCFF743FCW|/files/ENCFF743FCW/} processed by ChIP-seq ENCODE3 hg19 pipeline have a rescue ratio of 1.07 and a self consistency ratio of 2.03. According to ENCODE standards, having both rescue ratio and self consistency ratio values < 2 is recommended, but having only one of the ratio values < 2 is acceptable. audit_warning: Replicate concordance in ChIP-seq experiments is measured by calculating IDR values (Irreproducible Discovery Rate). ENCODE processed IDR thresholded peaks files {ENCFF360MGZ|/files/ENCFF360MGZ/}, {ENCFF448YLM|/files/ENCFF448YLM/} processed by ChIP-seq ENCODE3 hg19 pipeline have a rescue ratio of 1.07 and a self consistency ratio of 2.03. According to ENCODE standards, having both rescue ratio and self consistency ratio values < 2 is recommended, but having only one of the ratio values < 2 is acceptable. |
| Labels: |
|
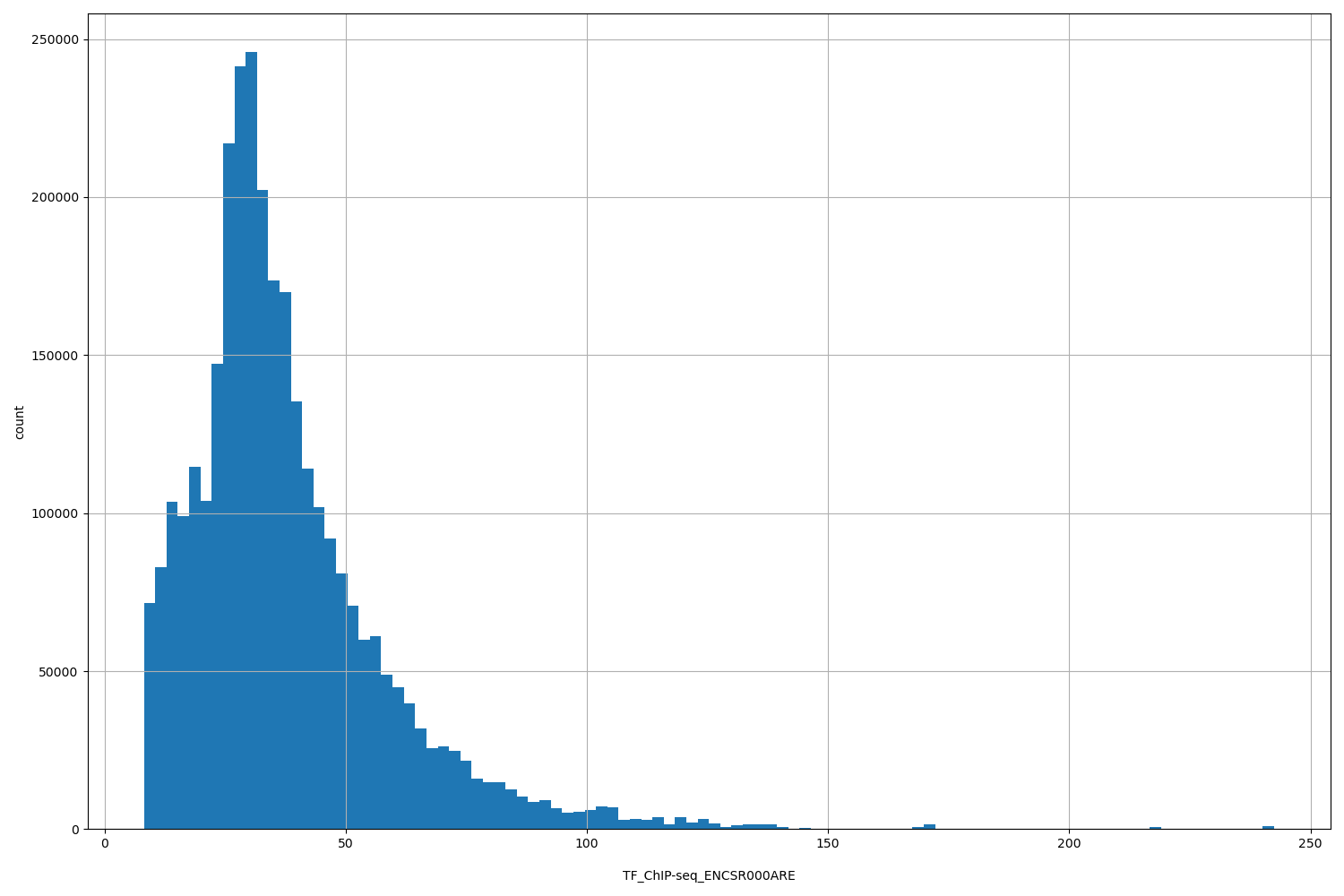
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| TF_ChIP-seq_ENCSR000ARE | float |
TF_ChIP-seq_ENCSR000ARE |
TF_ChIP-seq ENCSR000ARE [biosample_summary="Homo sapiens mammary epithelial cell female adult (50 years)" and target="EZH2"]
|
 |
[8.18, 242] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF743FCW.bed.gz | 140.69 KB | 3dfb5f076fde87b7896b22f33633d866 |
| ENCFF743FCW.bed.gz.dvc | 100.0 B | c8a8594a38930426fe3c6e67adac92e7 |
| ENCFF743FCW.tabix.bed.gz | 110.54 KB | adabc789272fb053ce82ff7aa053be8d |
| ENCFF743FCW.tabix.bed.gz.dvc | 106.0 B | 8dcaaef1c9660af89cb86f962b1ad507 |
| ENCFF743FCW.tabix.bed.gz.tbi | 35.8 KB | b28de683bcaab15a05e16cbf6fa7185d |
| ENCFF743FCW.tabix.bed.gz.tbi.dvc | 109.0 B | 43fbee90a55629cabea62a2d5b167746 |
| genomic_resource.yaml | 2.6 KB | 7016a8d67e5e61b452e0c2faf6d5716c |
| genomic_resource_original.yaml | 2.46 KB | 31866fabf2792af18d1a3c8e3344ba51 |
| statistics/ |