Histone_ChIP-seq_ENCSR960ZMM
| Id: | Histone_ChIP-seq/ENCSR960ZMM |
| Type: | position_score |
| Version: | 0 |
| Summary: |
HistoneChIP-seq ENCSR960ZMM [biosamplesummary="Homo sapiens OCI-LY1" and target="H4K20me1"] |
| Description: |
status: released biological_replicates: Rep 1, Rep 2 summary: output_type: replicated peaks audit_internal_action: Archived analysis {ENCAN902AXY|/analyses/ENCAN902AXY/} has in progress subobject quality standard {encode3-histone-chip|/quality-standards/encode3-histone-chip/} audit_not_compliant: Processed alignments file {ENCFF729YBU|/files/ENCFF729YBU/} processed by ChIP-seq ENCODE3 hg19 pipeline has 25188316 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting H4K20me1-human and investigated as a broad histone mark is 20 million usable fragments. The recommended value is > 45 million, but > 35 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/}) audit_warning: PBC1 (PCR Bottlenecking Coefficient 1, M1/M_distinct) is the ratio of the number of genomic locations where exactly one read maps uniquely (M1) to the number of genomic locations where some reads map (M_distinct). A PBC1 value in the range 0 - 0.5 is severe bottlenecking, 0.5 - 0.8 is moderate bottlenecking, 0.8 - 0.9 is mild bottlenecking, and > 0.9 is no bottlenecking. PBC1 value > 0.9 is recommended, but > 0.8 is acceptable. ENCODE processed alignments file {ENCFF454HDW|/files/ENCFF454HDW/} processed by ChIP-seq ENCODE3 hg19 pipeline was generated from a library with PBC1 value of 0.87. audit_warning: PBC2 (PCR Bottlenecking Coefficient 2, M1/M2) is the ratio of the number of genomic locations where exactly one read maps uniquely (M1) to the number of genomic locations where two reads map uniquely (M2). A PBC2 value in the range 0 - 1 is severe bottlenecking, 1 - 3 is moderate bottlenecking, 3 - 10 is mild bottlenecking, > 10 is no bottlenecking. PBC2 value > 10 is recommended, but > 3 is acceptable. ENCODE processed alignments file {ENCFF454HDW|/files/ENCFF454HDW/} processed by ChIP-seq ENCODE3 hg19 pipeline was generated from a library with PBC2 value of 7.65. |
| Labels: |
|
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| Histone_ChIP-seq_ENCSR960ZMM | float |
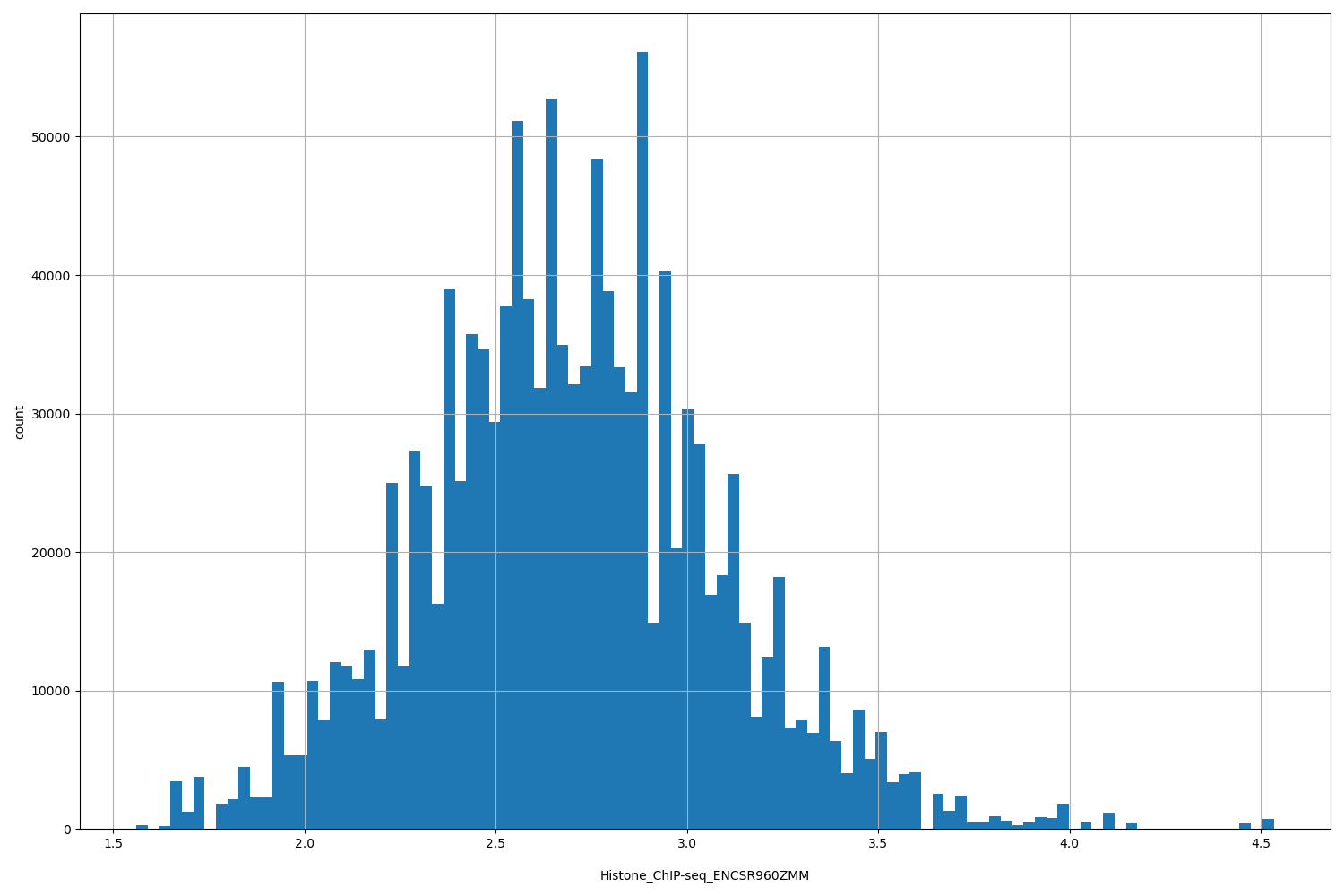
Histone_ChIP-seq_ENCSR960ZMM |
Histone_ChIP-seq ENCSR960ZMM [biosample_summary="Homo sapiens OCI-LY1" and target="H4K20me1"]
|
 |
[1.56, 4.53] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF521WMG.bed.gz | 78.63 KB | cdfbc158f7e7a2173f669b69c391bdb8 |
| ENCFF521WMG.bed.gz.dvc | 99.0 B | 920ef6b66a2e11511e0e218e6319177b |
| ENCFF521WMG.tabix.bed.gz | 40.74 KB | b0a2e348dfa9532326f54b6cd672f26d |
| ENCFF521WMG.tabix.bed.gz.dvc | 105.0 B | 846ffc3b08327a53efd3f3da115e854d |
| ENCFF521WMG.tabix.bed.gz.tbi | 31.18 KB | 9f555d1dd9e65eb74d167a6c859dbe59 |
| ENCFF521WMG.tabix.bed.gz.tbi.dvc | 109.0 B | 073fdfbcfa53f68f09d91f4debc28de4 |
| genomic_resource.yaml | 3.17 KB | b5abcd261995dbfa5d252ebba4fa3a0d |
| genomic_resource_original.yaml | 3.06 KB | 3255cb2c8b6035f98eb529954d3ef78e |
| statistics/ |