Histone_ChIP-seq_ENCSR948YYZ
| Id: | Histone_ChIP-seq/ENCSR948YYZ |
| Type: | position_score |
| Version: | 0 |
| Summary: |
HistoneChIP-seq ENCSR948YYZ [biosamplesummary="Homo sapiens gastrocnemius medialis tissue male adult (54 years)" and target="H3K27ac"] |
| Description: |
status: released biological_replicates: Rep 1, Rep 2 summary: male adult (54 years) output_type: replicated peaks audit_internal_action: Archived analysis {ENCAN658FEG|/analyses/ENCAN658FEG/} has in progress subobject quality standard {encode3-histone-chip|/quality-standards/encode3-histone-chip/} audit_warning: PBC1 (PCR Bottlenecking Coefficient 1, M1/M_distinct) is the ratio of the number of genomic locations where exactly one read maps uniquely (M1) to the number of genomic locations where some reads map (M_distinct). A PBC1 value in the range 0 - 0.5 is severe bottlenecking, 0.5 - 0.8 is moderate bottlenecking, 0.8 - 0.9 is mild bottlenecking, and > 0.9 is no bottlenecking. PBC1 value > 0.9 is recommended, but > 0.8 is acceptable. ENCODE processed alignments file {ENCFF258BZL|/files/ENCFF258BZL/} processed by ChIP-seq ENCODE3 GRCh38 pipeline was generated from a library with PBC1 value of 0.89. audit_warning: PBC2 (PCR Bottlenecking Coefficient 2, M1/M2) is the ratio of the number of genomic locations where exactly one read maps uniquely (M1) to the number of genomic locations where two reads map uniquely (M2). A PBC2 value in the range 0 - 1 is severe bottlenecking, 1 - 3 is moderate bottlenecking, 3 - 10 is mild bottlenecking, > 10 is no bottlenecking. PBC2 value > 10 is recommended, but > 3 is acceptable. ENCODE processed alignments file {ENCFF258BZL|/files/ENCFF258BZL/} processed by ChIP-seq ENCODE3 GRCh38 pipeline was generated from a library with PBC2 value of 9.58. |
| Labels: |
|
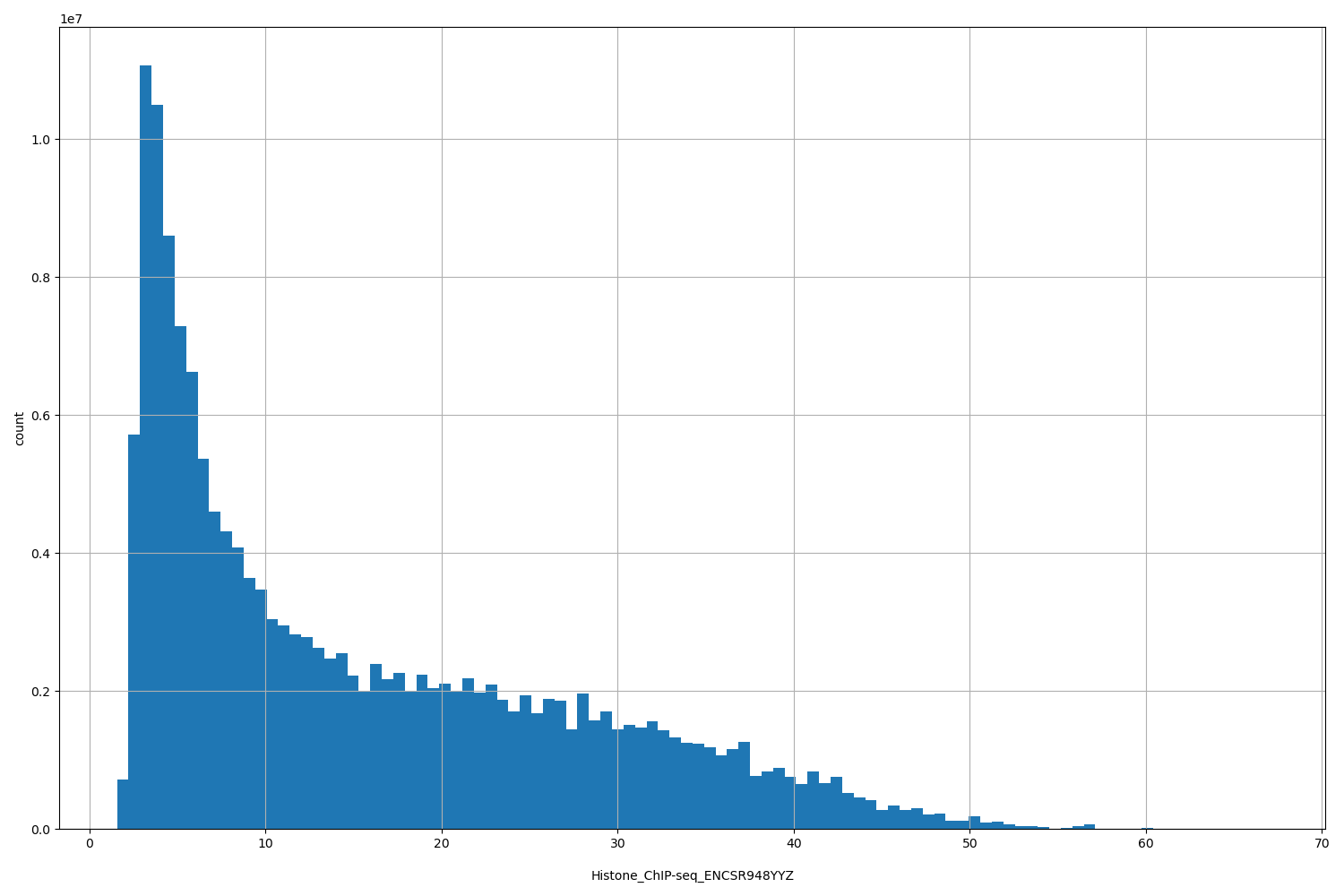
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| Histone_ChIP-seq_ENCSR948YYZ | float |
Histone_ChIP-seq_ENCSR948YYZ |
Histone_ChIP-seq ENCSR948YYZ [biosample_summary="Homo sapiens gastrocnemius medialis tissue male adult (54 years)" and target="H3K27ac"]
|
 |
[1.57, 66.9] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF369MQM.bed.gz | 2.1 MB | dd91ddf224cbf71e191f1fedcea24e8d |
| ENCFF369MQM.bed.gz.dvc | 101.0 B | ba0276fb41bfefe0caa95b77aa560250 |
| ENCFF369MQM.tabix.bed.gz | 918.46 KB | c95572c73c4aecdc11fb6a18111de05d |
| ENCFF369MQM.tabix.bed.gz.dvc | 106.0 B | 511f235dd16d68874496b790476d73fd |
| ENCFF369MQM.tabix.bed.gz.tbi | 222.17 KB | fb92b9321564313b05b325254da4b7c0 |
| ENCFF369MQM.tabix.bed.gz.tbi.dvc | 110.0 B | daa3e45c87114134cee9d0795ba7e630 |
| genomic_resource.yaml | 2.78 KB | 435e47ddf88493241b8288fa2dbffcb0 |
| genomic_resource_original.yaml | 2.63 KB | d894746319834c286a4563b2c8f3de98 |
| statistics/ |