Histone_ChIP-seq_ENCSR942NME
| Id: | Histone_ChIP-seq/ENCSR942NME |
| Type: | position_score |
| Version: | 0 |
| Summary: |
HistoneChIP-seq ENCSR942NME [biosamplesummary="Homo sapiens IMR-90" and target="H3K23ac"] |
| Description: |
status: released biological_replicates: Rep 1, Rep 2 summary: output_type: replicated peaks audit_internal_action: Archived analysis {ENCAN100MUZ|/analyses/ENCAN100MUZ/} has in progress subobject quality standard {encode3-histone-chip|/quality-standards/encode3-histone-chip/} audit_warning: Processed alignments file {ENCFF398ZPA|/files/ENCFF398ZPA/} processed by ChIP-seq ENCODE3 hg19 pipeline has 11756299 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting H3K23ac-human and investigated as a transcription factor is 10 million usable fragments. The recommended value is > 20 million, but > 10 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/} ) |
| Labels: |
|
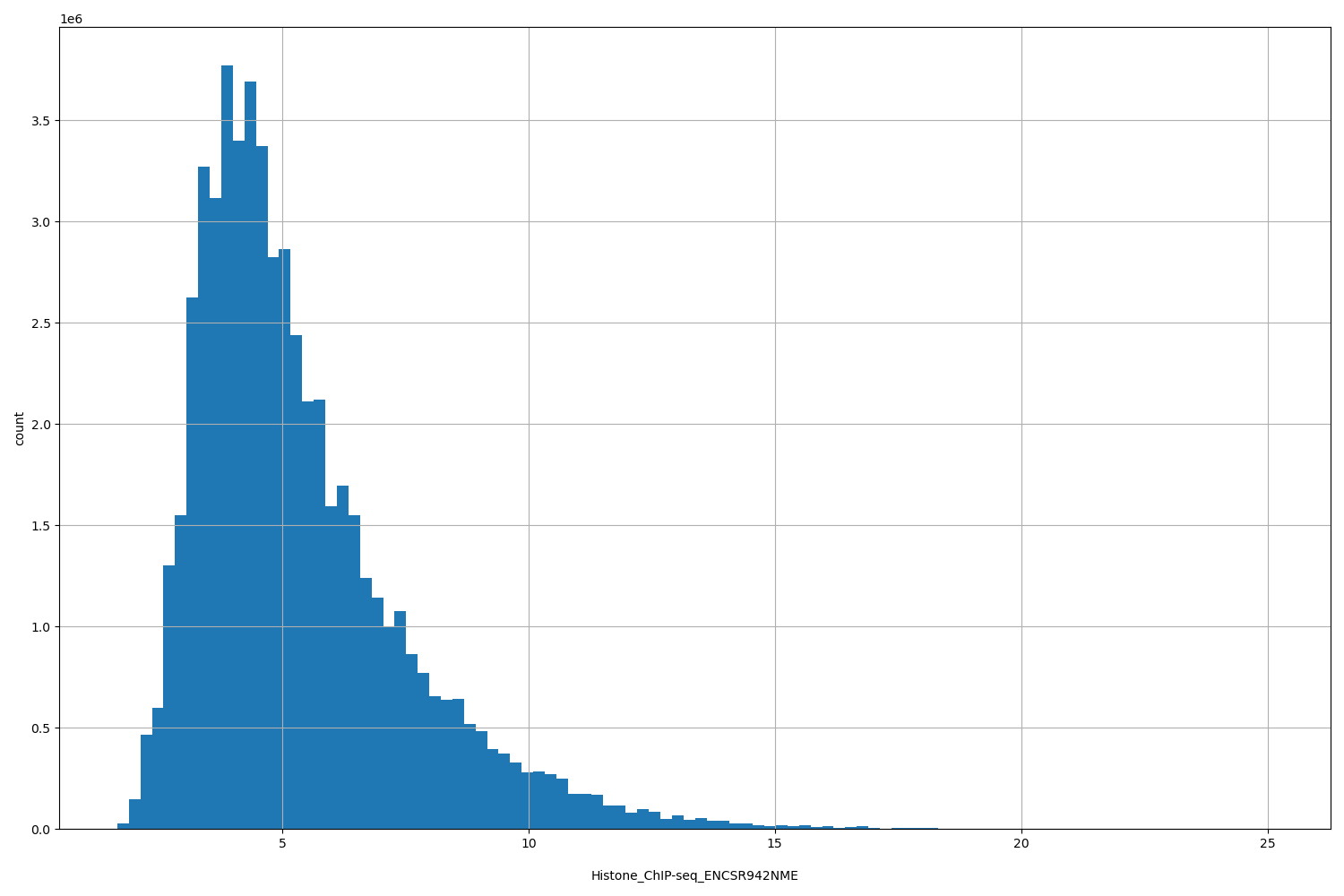
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| Histone_ChIP-seq_ENCSR942NME | float |
Histone_ChIP-seq_ENCSR942NME |
Histone_ChIP-seq ENCSR942NME [biosample_summary="Homo sapiens IMR-90" and target="H3K23ac"]
|
 |
[1.65, 25.1] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF712SFS.bed.gz | 3.43 MB | efa56569bedc545781316f68a83ea581 |
| ENCFF712SFS.bed.gz.dvc | 101.0 B | 6086968b533d61e559039b6e192fda13 |
| ENCFF712SFS.tabix.bed.gz | 1.54 MB | 292f5e2c2043704153ad779878e6a336 |
| ENCFF712SFS.tabix.bed.gz.dvc | 107.0 B | a48355d21597ce255ecbff9aa3fd4061 |
| ENCFF712SFS.tabix.bed.gz.tbi | 320.87 KB | 369db9e4cd5b2d83a233fd77c605e568 |
| ENCFF712SFS.tabix.bed.gz.tbi.dvc | 110.0 B | d6c861f306d0a8b701b0121e34342e9a |
| genomic_resource.yaml | 1.86 KB | dee7a512976fc16dbf6d9a04b9ef031b |
| genomic_resource_original.yaml | 1.75 KB | 2d95296551c0e822f5cb37a7f0adbb7b |
| statistics/ |