Histone_ChIP-seq_ENCSR920YBY
| Id: | Histone_ChIP-seq/ENCSR920YBY |
| Type: | position_score |
| Version: | 0 |
| Summary: |
HistoneChIP-seq ENCSR920YBY [biosamplesummary="Homo sapiens common myeloid progenitor, CD34-positive" and target="H3K36me3"] |
| Description: |
status: released biological_replicates: Rep 1, Rep 2 summary: output_type: pseudoreplicated peaks audit_internal_action: Released analysis {ENCAN741RPX|/analyses/ENCAN741RPX/} has in progress subobject quality standard {encode4-histone-chip|/quality-standards/encode4-histone-chip/} audit_internal_action: Released analysis {ENCAN741RPX|/analyses/ENCAN741RPX/} has in progress subobject document {91add1c2-64fb-4d16-929c-0a16a25dac88|/documents/91add1c2-64fb-4d16-929c-0a16a25dac88/} audit_not_compliant: Processed alignments file {ENCFF641LRJ|/files/ENCFF641LRJ/} processed by ChIP-seq ENCODE4 v1.6.1 GRCh38 pipeline has 9254208 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting H3K36me3-human and investigated as a broad histone mark is 20 million usable fragments. The recommended value is > 45 million, but > 35 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/}) audit_not_compliant: Processed alignments file {ENCFF325EXU|/files/ENCFF325EXU/} processed by ChIP-seq ENCODE4 v1.6.1 GRCh38 pipeline has 21627399 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting H3K36me3-human and investigated as a broad histone mark is 20 million usable fragments. The recommended value is > 45 million, but > 35 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/}) |
| Labels: |
|
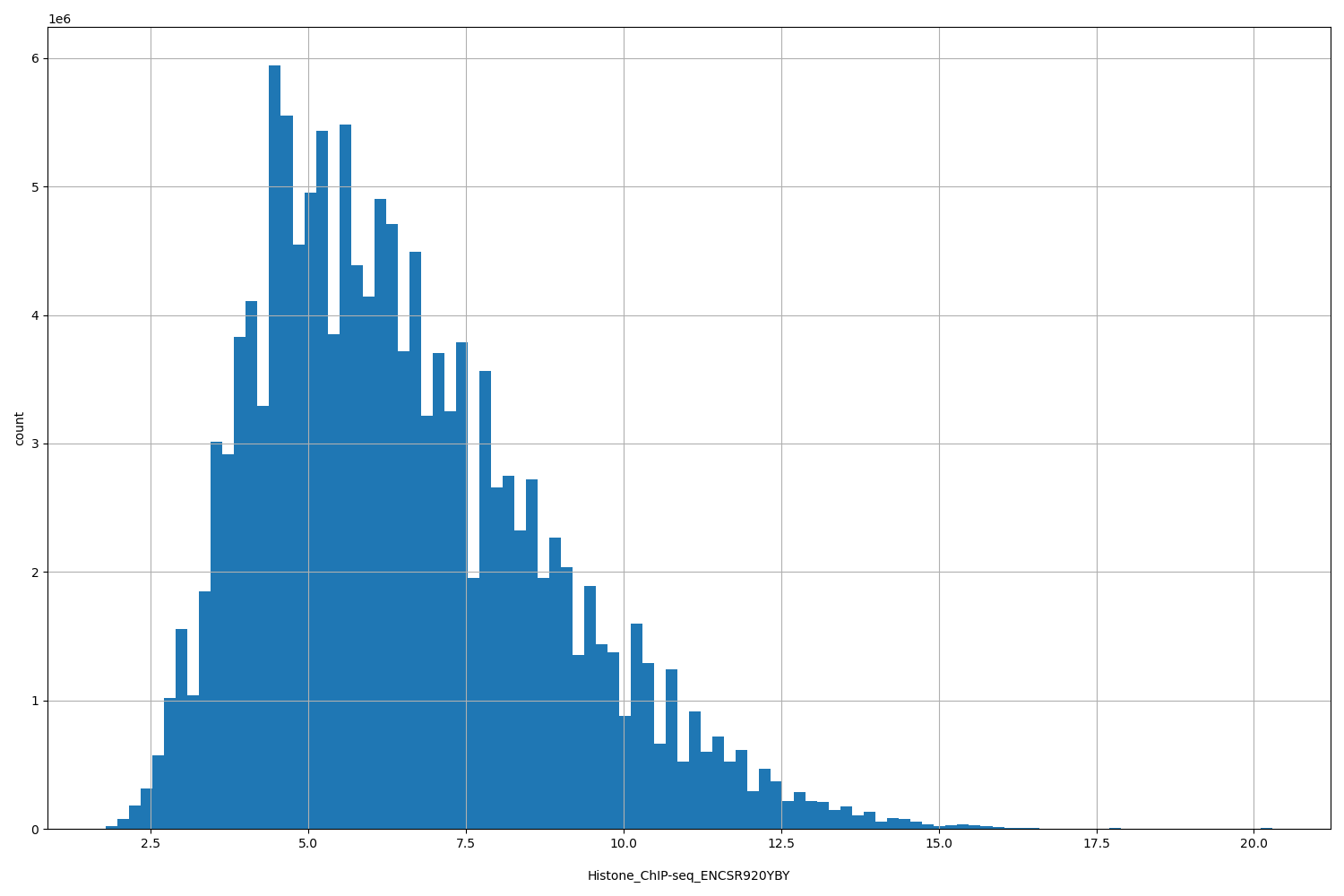
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| Histone_ChIP-seq_ENCSR920YBY | float |
Histone_ChIP-seq_ENCSR920YBY |
Histone_ChIP-seq ENCSR920YBY [biosample_summary="Homo sapiens common myeloid progenitor, CD34-positive" and target="H3K36me3"]
|
 |
[1.79, 20.3] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF996OPQ.bed.gz | 3.54 MB | 6cf971968654345d340062a83315346f |
| ENCFF996OPQ.bed.gz.dvc | 101.0 B | 68ce46bc3d0568e520131543d1ff7d10 |
| ENCFF996OPQ.tabix.bed.gz | 1.68 MB | 72f89784866a524903f8d407825ccb83 |
| ENCFF996OPQ.tabix.bed.gz.dvc | 107.0 B | d70226d7b7f54995d85226b7bbf6ce40 |
| ENCFF996OPQ.tabix.bed.gz.tbi | 170.4 KB | f0fcd2b1a21f1d0ef0fd261e6ec15e88 |
| ENCFF996OPQ.tabix.bed.gz.tbi.dvc | 110.0 B | 701ba662f280934f40915f24247b9124 |
| genomic_resource.yaml | 2.79 KB | d04c9f3fb55c00ccc1f4fda902c93979 |
| genomic_resource_original.yaml | 2.65 KB | 016eaf0991e5841a676149846a286426 |
| statistics/ |