Histone_ChIP-seq_ENCSR843KHS
| Id: | Histone_ChIP-seq/ENCSR843KHS |
| Type: | position_score |
| Version: | 0 |
| Summary: |
HistoneChIP-seq ENCSR843KHS [biosamplesummary="Homo sapiens psoas muscle tissue male child (3 years)" and target="H3K27me3"] |
| Description: |
status: released biological_replicates: Rep 1 summary: male child (3 years) output_type: pseudoreplicated peaks audit_internal_action: Archived analysis {ENCAN520INO|/analyses/ENCAN520INO/} has in progress subobject quality standard {encode3-histone-chip|/quality-standards/encode3-histone-chip/} audit_not_compliant: Processed alignments file {ENCFF768PBV|/files/ENCFF768PBV/} processed by ChIP-seq ENCODE3 hg19 pipeline has 21161513 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting H3K27me3-human and investigated as a broad histone mark is 20 million usable fragments. The recommended value is > 45 million, but > 35 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/}) audit_warning: PBC1 (PCR Bottlenecking Coefficient 1, M1/M_distinct) is the ratio of the number of genomic locations where exactly one read maps uniquely (M1) to the number of genomic locations where some reads map (M_distinct). A PBC1 value in the range 0 - 0.5 is severe bottlenecking, 0.5 - 0.8 is moderate bottlenecking, 0.8 - 0.9 is mild bottlenecking, and > 0.9 is no bottlenecking. PBC1 value > 0.9 is recommended, but > 0.8 is acceptable. ENCODE processed alignments file {ENCFF768PBV|/files/ENCFF768PBV/} processed by ChIP-seq ENCODE3 hg19 pipeline was generated from a library with PBC1 value of 0.90. audit_warning: PBC2 (PCR Bottlenecking Coefficient 2, M1/M2) is the ratio of the number of genomic locations where exactly one read maps uniquely (M1) to the number of genomic locations where two reads map uniquely (M2). A PBC2 value in the range 0 - 1 is severe bottlenecking, 1 - 3 is moderate bottlenecking, 3 - 10 is mild bottlenecking, > 10 is no bottlenecking. PBC2 value > 10 is recommended, but > 3 is acceptable. ENCODE processed alignments file {ENCFF768PBV|/files/ENCFF768PBV/} processed by ChIP-seq ENCODE3 hg19 pipeline was generated from a library with PBC2 value of 9.65. |
| Labels: |
|
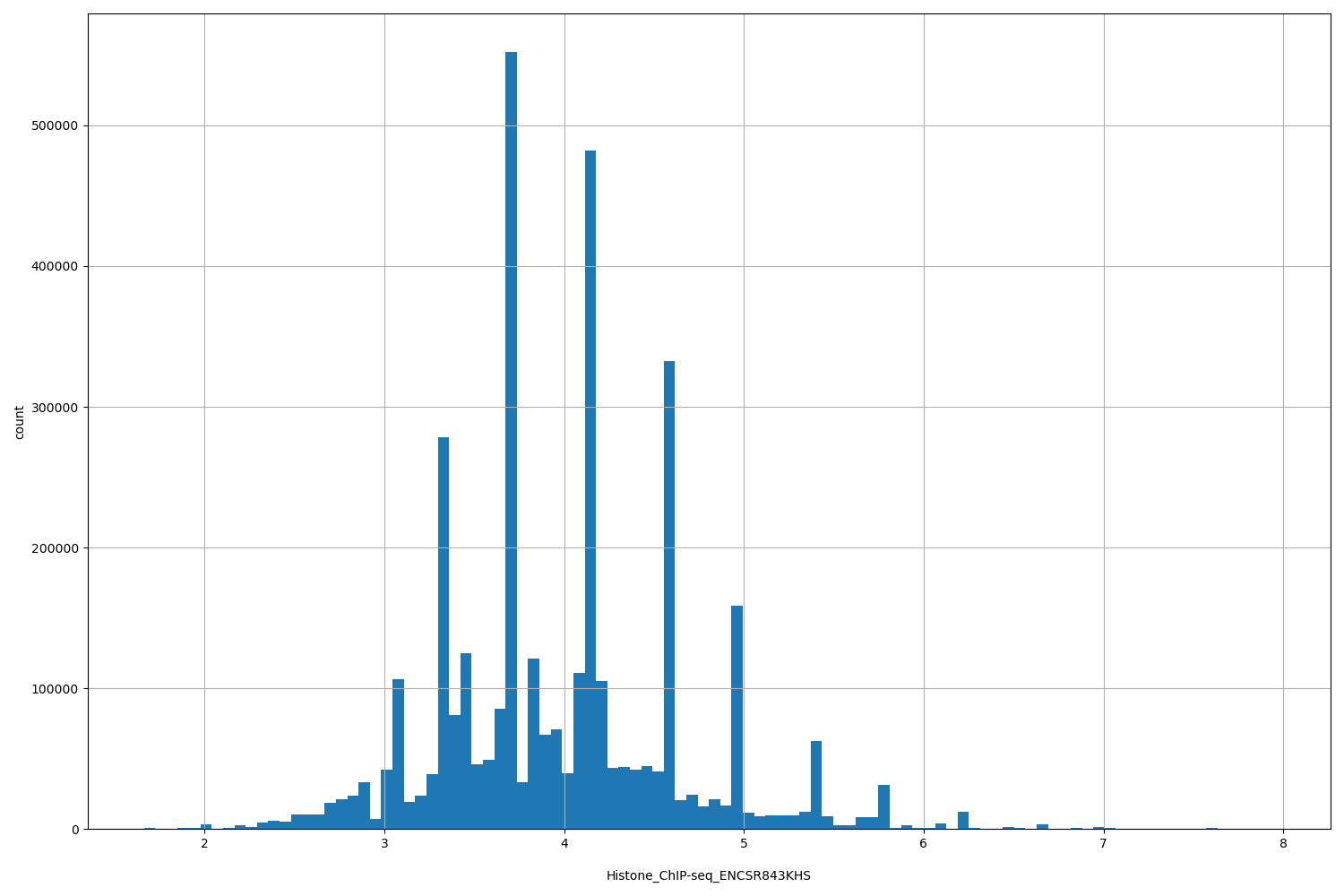
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| Histone_ChIP-seq_ENCSR843KHS | float |
Histone_ChIP-seq_ENCSR843KHS |
Histone_ChIP-seq ENCSR843KHS [biosample_summary="Homo sapiens psoas muscle tissue male child (3 years)" and target="H3K27me3"]
|
 |
[1.66, 7.95] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF006QNM.bed.gz | 208.63 KB | 134e4a5f1d4eb3b931f1ab7d4de54000 |
| ENCFF006QNM.bed.gz.dvc | 100.0 B | 688000ececbefbe0fa19a319dbf150d3 |
| ENCFF006QNM.tabix.bed.gz | 109.7 KB | 642e9694e9e5fef265a9ff7907d60e9c |
| ENCFF006QNM.tabix.bed.gz.dvc | 106.0 B | cdc5c9e76bd6466eec2c7a4955dd5f6b |
| ENCFF006QNM.tabix.bed.gz.tbi | 85.21 KB | 7da007fd4932eb32f11ba7fdce5e3821 |
| ENCFF006QNM.tabix.bed.gz.tbi.dvc | 109.0 B | 5f9efdb12eb639b4b0d7d348080b177f |
| genomic_resource.yaml | 3.23 KB | 9c77986935f664424ed72bd39b15c7fc |
| genomic_resource_original.yaml | 3.09 KB | 5d2b99cd0bb68c9a0f169e812f45be3b |
| statistics/ |