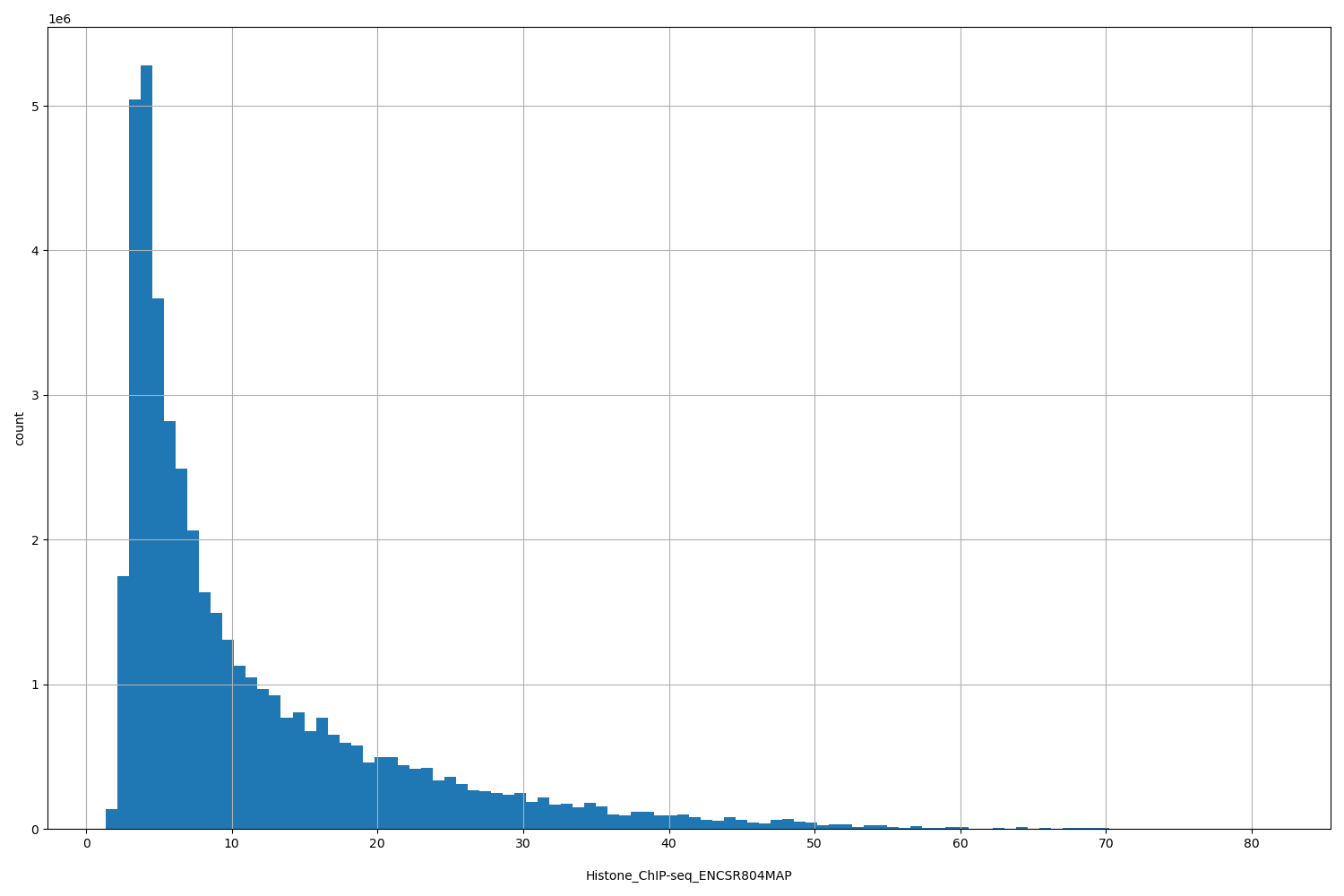
Histone_ChIP-seq_ENCSR804MAP
| Id: | Histone_ChIP-seq/ENCSR804MAP |
| Type: | position_score |
| Version: | 0 |
| Summary: |
HistoneChIP-seq ENCSR804MAP [biosamplesummary="Homo sapiens HUES64" and target="H3K27ac"] |
| Description: |
status: released biological_replicates: Rep 1, Rep 2 summary: output_type: replicated peaks audit_internal_action: Archived analysis {ENCAN102CQK|/analyses/ENCAN102CQK/} has in progress subobject quality standard {encode3-histone-chip|/quality-standards/encode3-histone-chip/} audit_warning: Processed redacted alignments file {ENCFF846LDC|/files/ENCFF846LDC/} processed by ChIP-seq ENCODE3 hg19 pipeline has 18723635 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting H3K27ac-human and investigated as a narrow histone mark is 10 million usable fragments. The recommended value is > 20 million, but > 10 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/}) |
| Labels: |
|
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| Histone_ChIP-seq_ENCSR804MAP | float |
Histone_ChIP-seq_ENCSR804MAP |
Histone_ChIP-seq ENCSR804MAP [biosample_summary="Homo sapiens HUES64" and target="H3K27ac"]
|
 |
[1.36, 81.4] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF196TWN.bed.gz | 1.55 MB | efd5fa50421afa52d5badcb383c8bd71 |
| ENCFF196TWN.bed.gz.dvc | 101.0 B | dd724bf6a5c4b4a6f129a0c980c1be91 |
| ENCFF196TWN.tabix.bed.gz | 708.01 KB | 6796433f713efb8d514de3b2867565dd |
| ENCFF196TWN.tabix.bed.gz.dvc | 106.0 B | 5f9643f34d1e15dfd0d72c3fd944951f |
| ENCFF196TWN.tabix.bed.gz.tbi | 271.48 KB | c013e8170ed0196ee9510b7f79a10d60 |
| ENCFF196TWN.tabix.bed.gz.tbi.dvc | 110.0 B | f8f9b103a1f97bf8102dfd3d89ca667c |
| genomic_resource.yaml | 1.87 KB | 73bee7c2805f058060fc2a4678236b65 |
| genomic_resource_original.yaml | 1.76 KB | 06f0925de6413cd1ebcb551355a02620 |
| statistics/ |