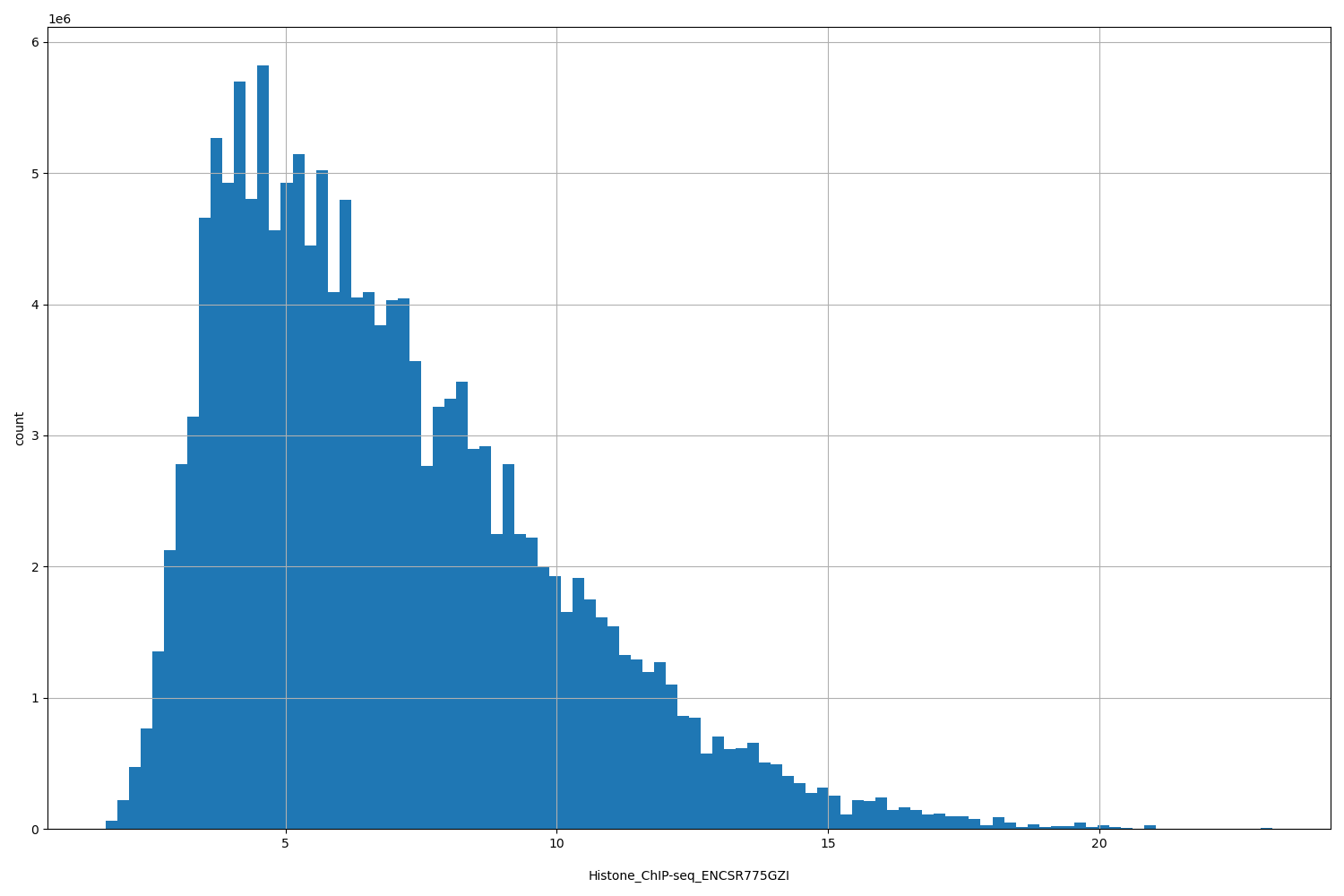
Histone_ChIP-seq_ENCSR775GZI
| Id: | Histone_ChIP-seq/ENCSR775GZI |
| Type: | position_score |
| Version: | 0 |
| Summary: |
HistoneChIP-seq ENCSR775GZI [biosamplesummary="Homo sapiens HUES48" and target="H3K36me3"] |
| Description: |
status: released biological_replicates: Rep 1, Rep 2 summary: output_type: pseudoreplicated peaks audit_internal_action: Released analysis {ENCAN700VCV|/analyses/ENCAN700VCV/} has in progress subobject quality standard {encode4-histone-chip|/quality-standards/encode4-histone-chip/} audit_internal_action: Released analysis {ENCAN700VCV|/analyses/ENCAN700VCV/} has in progress subobject document {d8c7b396-b0d3-4e5b-9a6b-e1bac7bf80c0|/documents/d8c7b396-b0d3-4e5b-9a6b-e1bac7bf80c0/} audit_not_compliant: Processed alignments file {ENCFF822VVN|/files/ENCFF822VVN/} processed by ChIP-seq ENCODE4 v1.6.1 GRCh38 pipeline has 30926287 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting H3K36me3-human and investigated as a broad histone mark is 20 million usable fragments. The recommended value is > 45 million, but > 35 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/}) audit_not_compliant: Processed alignments file {ENCFF022AIU|/files/ENCFF022AIU/} processed by ChIP-seq ENCODE4 v1.6.1 GRCh38 pipeline has 21653155 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting H3K36me3-human and investigated as a broad histone mark is 20 million usable fragments. The recommended value is > 45 million, but > 35 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/}) |
| Labels: |
|
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| Histone_ChIP-seq_ENCSR775GZI | float |
Histone_ChIP-seq_ENCSR775GZI |
Histone_ChIP-seq ENCSR775GZI [biosample_summary="Homo sapiens HUES48" and target="H3K36me3"]
|
 |
[1.69, 23.2] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF716JGC.bed.gz | 2.7 MB | 85bf5325cb90316ffa2d8f639c8adf25 |
| ENCFF716JGC.bed.gz.dvc | 101.0 B | 5e96e9c7ca47bb98786a8e2854221911 |
| ENCFF716JGC.tabix.bed.gz | 1.26 MB | b6a942bbee010e965ff6f0e9b90730a9 |
| ENCFF716JGC.tabix.bed.gz.dvc | 107.0 B | e940840652726e179ed6920fd4c2eb17 |
| ENCFF716JGC.tabix.bed.gz.tbi | 156.58 KB | c9ff4593fa99830626f63412bd2eaf43 |
| ENCFF716JGC.tabix.bed.gz.tbi.dvc | 110.0 B | c8c7d7f8f38188281a50446207d14bca |
| genomic_resource.yaml | 2.61 KB | 008e1dc83d4004395c252b8ed6a0657f |
| genomic_resource_original.yaml | 2.5 KB | 0495e132956f100f2fbeb012aefe9b77 |
| statistics/ |