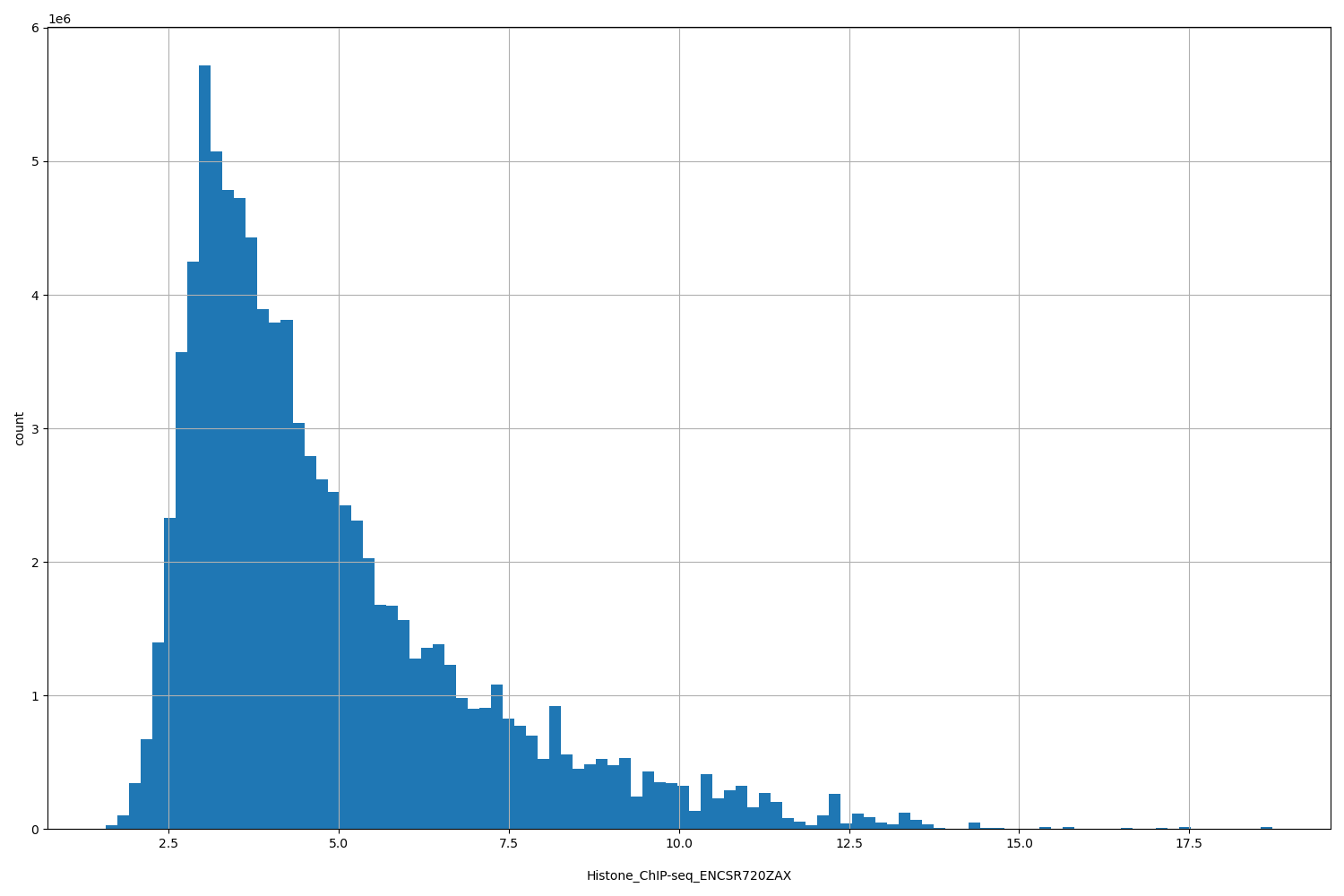
Histone_ChIP-seq_ENCSR720ZAX
| Id: | Histone_ChIP-seq/ENCSR720ZAX |
| Type: | position_score |
| Version: | 0 |
| Summary: |
HistoneChIP-seq ENCSR720ZAX [biosamplesummary="Homo sapiens neural progenitor cell originated from H9" and target="H4K20me1"] |
| Description: |
status: released biological_replicates: Rep 1, Rep 2 summary: female embryo (5 days) output_type: pseudoreplicated peaks audit_internal_action: Released analysis {ENCAN853NVM|/analyses/ENCAN853NVM/} has in progress subobject quality standard {encode4-histone-chip|/quality-standards/encode4-histone-chip/} audit_internal_action: Released analysis {ENCAN853NVM|/analyses/ENCAN853NVM/} has in progress subobject document {9391a777-5de7-40f1-9efb-6b2ee2fb5ccf|/documents/9391a777-5de7-40f1-9efb-6b2ee2fb5ccf/} audit_warning: Processed alignments file {ENCFF621GTK|/files/ENCFF621GTK/} processed by ChIP-seq ENCODE4 v1.6.1 GRCh38 pipeline has 40685555 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting H4K20me1-human and investigated as a broad histone mark is 20 million usable fragments. The recommended value is > 45 million, but > 35 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/}) audit_warning: Processed alignments file {ENCFF100RPU|/files/ENCFF100RPU/} processed by ChIP-seq ENCODE4 v1.6.1 GRCh38 pipeline has 40827867 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting H4K20me1-human and investigated as a broad histone mark is 20 million usable fragments. The recommended value is > 45 million, but > 35 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/}) |
| Labels: |
|
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| Histone_ChIP-seq_ENCSR720ZAX | float |
Histone_ChIP-seq_ENCSR720ZAX |
Histone_ChIP-seq ENCSR720ZAX [biosample_summary="Homo sapiens neural progenitor cell originated from H9" and target="H4K20me1"]
|
 |
[1.58, 18.7] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF603XUN.bed.gz | 1.51 MB | 7f10dbd07e98ad0b7ba18da4c5b0b747 |
| ENCFF603XUN.bed.gz.dvc | 101.0 B | c50f2225044ef12a0a8e48d1064ba9ff |
| ENCFF603XUN.tabix.bed.gz | 716.76 KB | 4e6cab986037080f11f12785192f572e |
| ENCFF603XUN.tabix.bed.gz.dvc | 106.0 B | dfa94b8dd35f179beaa9a9822526223d |
| ENCFF603XUN.tabix.bed.gz.tbi | 157.9 KB | 228748987836df20b41fe5b793bd02bb |
| ENCFF603XUN.tabix.bed.gz.tbi.dvc | 110.0 B | cd87932885add314372fc0bcd7256d04 |
| genomic_resource.yaml | 2.71 KB | 8b7847374d0e4528fd13baee895dfcc7 |
| genomic_resource_original.yaml | 2.57 KB | 1cdfac54049cc824d937bc6fb86a4bac |
| statistics/ |