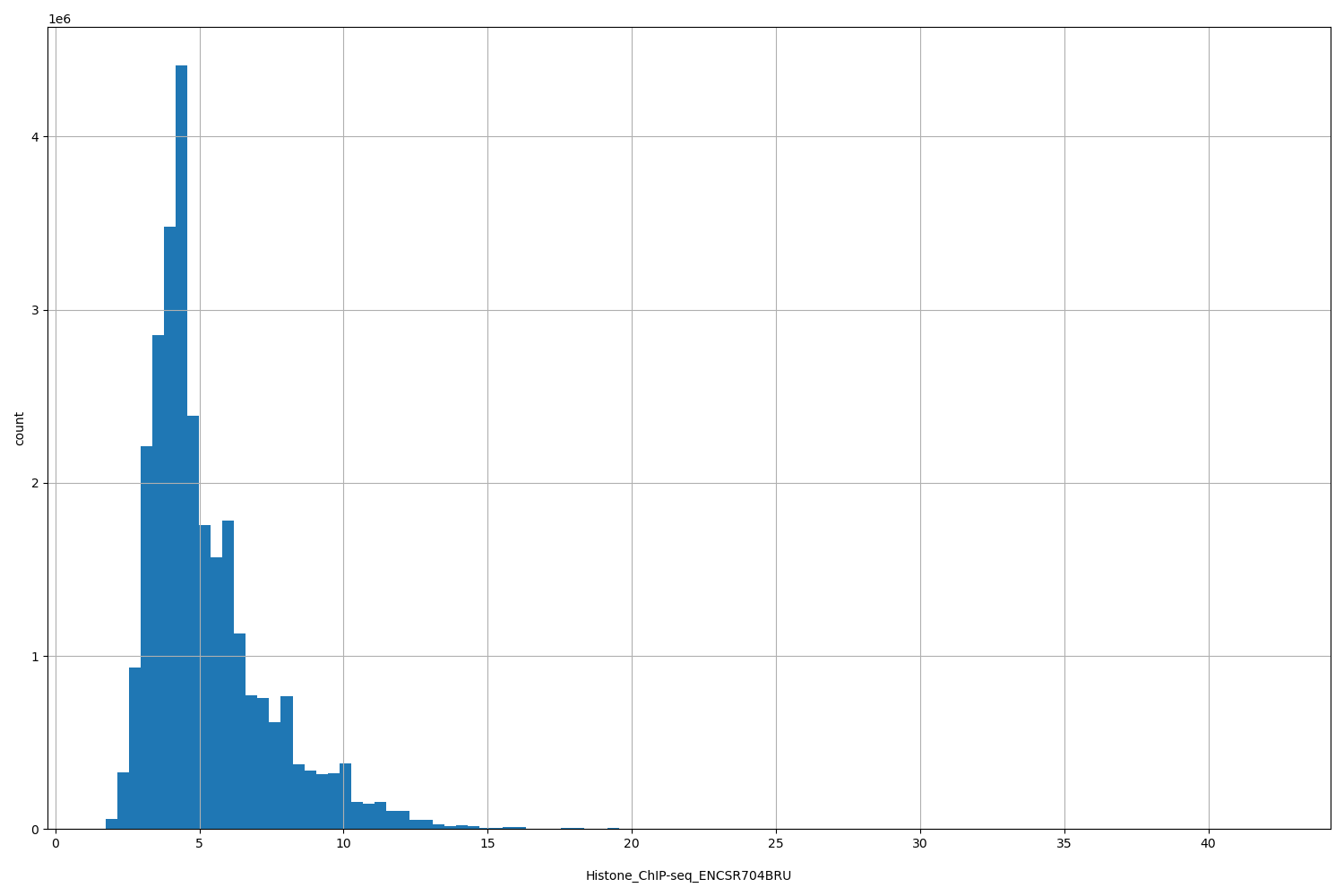
Histone_ChIP-seq_ENCSR704BRU
| Id: | Histone_ChIP-seq/ENCSR704BRU |
| Type: | position_score |
| Version: | 0 |
| Summary: |
HistoneChIP-seq ENCSR704BRU [biosamplesummary="Homo sapiens iPS DF 19.11" and target="H3K9me3"] |
| Description: |
status: released biological_replicates: Rep 1, Rep 2 summary: output_type: pseudoreplicated peaks audit_internal_action: Released analysis {ENCAN145JPG|/analyses/ENCAN145JPG/} has in progress subobject quality standard {encode4-histone-chip|/quality-standards/encode4-histone-chip/} audit_internal_action: Released analysis {ENCAN145JPG|/analyses/ENCAN145JPG/} has in progress subobject document {6e0cd75d-18a1-4348-a8dc-ccd08d3b25c5|/documents/6e0cd75d-18a1-4348-a8dc-ccd08d3b25c5/} audit_not_compliant: Processed unfiltered alignments file {ENCFF031STP|/files/ENCFF031STP/} processed by ChIP-seq ENCODE4 v1.6.1 GRCh38 pipeline has 28561796 mapped reads. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting H3K9me3-human and investigated as a broad histone mark is 35 million mapped reads. The recommended value is > 45 million, but > 35 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/}) audit_not_compliant: Processed unfiltered alignments file {ENCFF438TGC|/files/ENCFF438TGC/} processed by ChIP-seq ENCODE4 v1.6.1 GRCh38 pipeline has 27788090 mapped reads. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting H3K9me3-human and investigated as a broad histone mark is 35 million mapped reads. The recommended value is > 45 million, but > 35 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/}) |
| Labels: |
|
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| Histone_ChIP-seq_ENCSR704BRU | float |
Histone_ChIP-seq_ENCSR704BRU |
Histone_ChIP-seq ENCSR704BRU [biosample_summary="Homo sapiens iPS DF 19.11" and target="H3K9me3"]
|
 |
[1.76, 42.2] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF839TYU.bed.gz | 1.78 MB | 776be5d2eeb8aee36ee81ac81a8f85bb |
| ENCFF839TYU.bed.gz.dvc | 101.0 B | a181eb1f086b8b097965e3a1f9d82ef0 |
| ENCFF839TYU.tabix.bed.gz | 954.39 KB | 63c3ecc1dbd3195db7f87da7475bff25 |
| ENCFF839TYU.tabix.bed.gz.dvc | 106.0 B | 896663496d62514bcae441dc11d1978b |
| ENCFF839TYU.tabix.bed.gz.tbi | 346.73 KB | dbe3e29b0c4ab140492f4a8a27cb9b5b |
| ENCFF839TYU.tabix.bed.gz.tbi.dvc | 110.0 B | eff6f26142c10f7bc39ae53a5d3bbc5d |
| genomic_resource.yaml | 2.63 KB | 2ba1eee4f9e955bd638e4c5140b8c353 |
| genomic_resource_original.yaml | 2.52 KB | 6cfc18ac802ac5f0dbf03b0603058c97 |
| statistics/ |