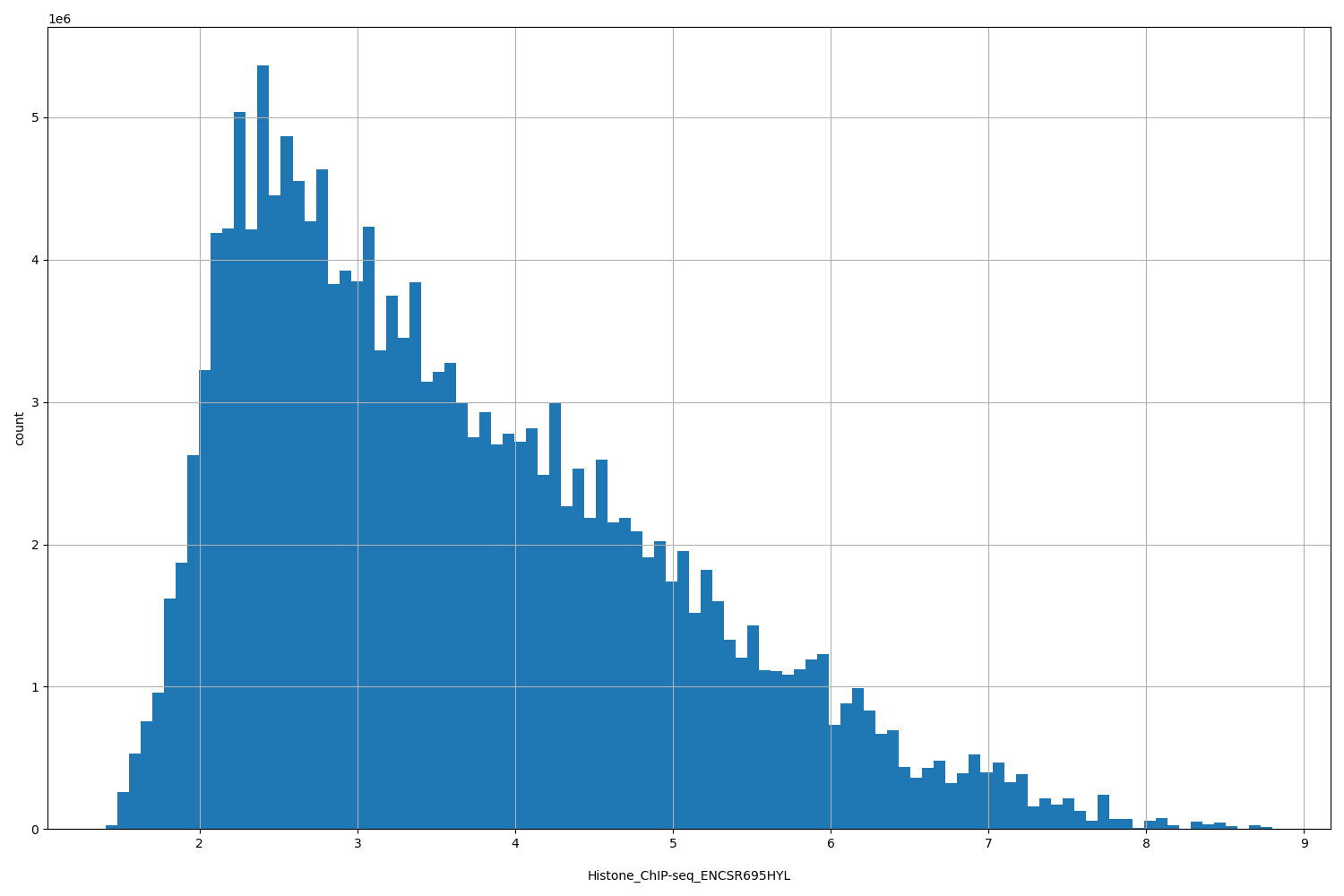
Histone_ChIP-seq_ENCSR695HYL
| Id: | Histone_ChIP-seq/ENCSR695HYL |
| Type: | position_score |
| Version: | 0 |
| Summary: |
HistoneChIP-seq ENCSR695HYL [biosamplesummary="Homo sapiens hepatocyte originated from H9" and target="H3K79me2"] |
| Description: |
status: released biological_replicates: Rep 1, Rep 2 summary: female embryo (5 days) output_type: replicated peaks audit_internal_action: Archived analysis {ENCAN601EBJ|/analyses/ENCAN601EBJ/} has in progress subobject quality standard {encode3-histone-chip|/quality-standards/encode3-histone-chip/} audit_warning: PBC1 (PCR Bottlenecking Coefficient 1, M1/M_distinct) is the ratio of the number of genomic locations where exactly one read maps uniquely (M1) to the number of genomic locations where some reads map (M_distinct). A PBC1 value in the range 0 - 0.5 is severe bottlenecking, 0.5 - 0.8 is moderate bottlenecking, 0.8 - 0.9 is mild bottlenecking, and > 0.9 is no bottlenecking. PBC1 value > 0.9 is recommended, but > 0.8 is acceptable. ENCODE processed alignments file {ENCFF764HMA|/files/ENCFF764HMA/} processed by ChIP-seq ENCODE3 hg19 pipeline was generated from a library with PBC1 value of 0.89. audit_warning: PBC2 (PCR Bottlenecking Coefficient 2, M1/M2) is the ratio of the number of genomic locations where exactly one read maps uniquely (M1) to the number of genomic locations where two reads map uniquely (M2). A PBC2 value in the range 0 - 1 is severe bottlenecking, 1 - 3 is moderate bottlenecking, 3 - 10 is mild bottlenecking, > 10 is no bottlenecking. PBC2 value > 10 is recommended, but > 3 is acceptable. ENCODE processed alignments file {ENCFF764HMA|/files/ENCFF764HMA/} processed by ChIP-seq ENCODE3 hg19 pipeline was generated from a library with PBC2 value of 8.93. |
| Labels: |
|
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| Histone_ChIP-seq_ENCSR695HYL | float |
Histone_ChIP-seq_ENCSR695HYL |
Histone_ChIP-seq ENCSR695HYL [biosample_summary="Homo sapiens hepatocyte originated from H9" and target="H3K79me2"]
|
 |
[1.41, 8.8] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF999FNX.bed.gz | 2.21 MB | efdc4bbe4332f33bd20dfa6b5b5069c3 |
| ENCFF999FNX.bed.gz.dvc | 101.0 B | 075b5400c3438dd1c0d01833cf4bb5a4 |
| ENCFF999FNX.tabix.bed.gz | 1007.67 KB | 9bf4b90b73fc973da1ac163b84017691 |
| ENCFF999FNX.tabix.bed.gz.dvc | 107.0 B | 2fe18a4fc9ad95383ec80e39149730a0 |
| ENCFF999FNX.tabix.bed.gz.tbi | 139.51 KB | 1916caa41f564ba6a5a33a86734674eb |
| ENCFF999FNX.tabix.bed.gz.tbi.dvc | 110.0 B | df322c374e2793ccd07e721d2a87b214 |
| genomic_resource.yaml | 2.74 KB | 204d4e4398c09d9f84c09eb25f440780 |
| genomic_resource_original.yaml | 2.61 KB | 32e1fcfc4c01eace6416d7922235b362 |
| statistics/ |