Histone_ChIP-seq_ENCSR692CTK
| Id: | Histone_ChIP-seq/ENCSR692CTK |
| Type: | position_score |
| Version: | 0 |
| Summary: |
HistoneChIP-seq ENCSR692CTK [biosamplesummary="Homo sapiens neuronal stem cell originated from H1" and target="H3K27me3"] |
| Description: |
status: released biological_replicates: Rep 1, Rep 2 summary: male embryo output_type: pseudoreplicated peaks audit_internal_action: Released analysis {ENCAN551COK|/analyses/ENCAN551COK/} has in progress subobject quality standard {encode4-histone-chip|/quality-standards/encode4-histone-chip/} audit_internal_action: Released analysis {ENCAN551COK|/analyses/ENCAN551COK/} has in progress subobject document {cdf43670-c1fb-46e4-86ab-26f4aac5f40d|/documents/cdf43670-c1fb-46e4-86ab-26f4aac5f40d/} audit_not_compliant: Processed alignments file {ENCFF972LPQ|/files/ENCFF972LPQ/} processed by ChIP-seq ENCODE4 v1.6.1 GRCh38 pipeline has 28529417 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting H3K27me3-human and investigated as a broad histone mark is 20 million usable fragments. The recommended value is > 45 million, but > 35 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/}) audit_warning: Processed alignments file {ENCFF418RUM|/files/ENCFF418RUM/} processed by ChIP-seq ENCODE4 v1.6.1 GRCh38 pipeline has 36010117 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting H3K27me3-human and investigated as a broad histone mark is 20 million usable fragments. The recommended value is > 45 million, but > 35 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/}) |
| Labels: |
|
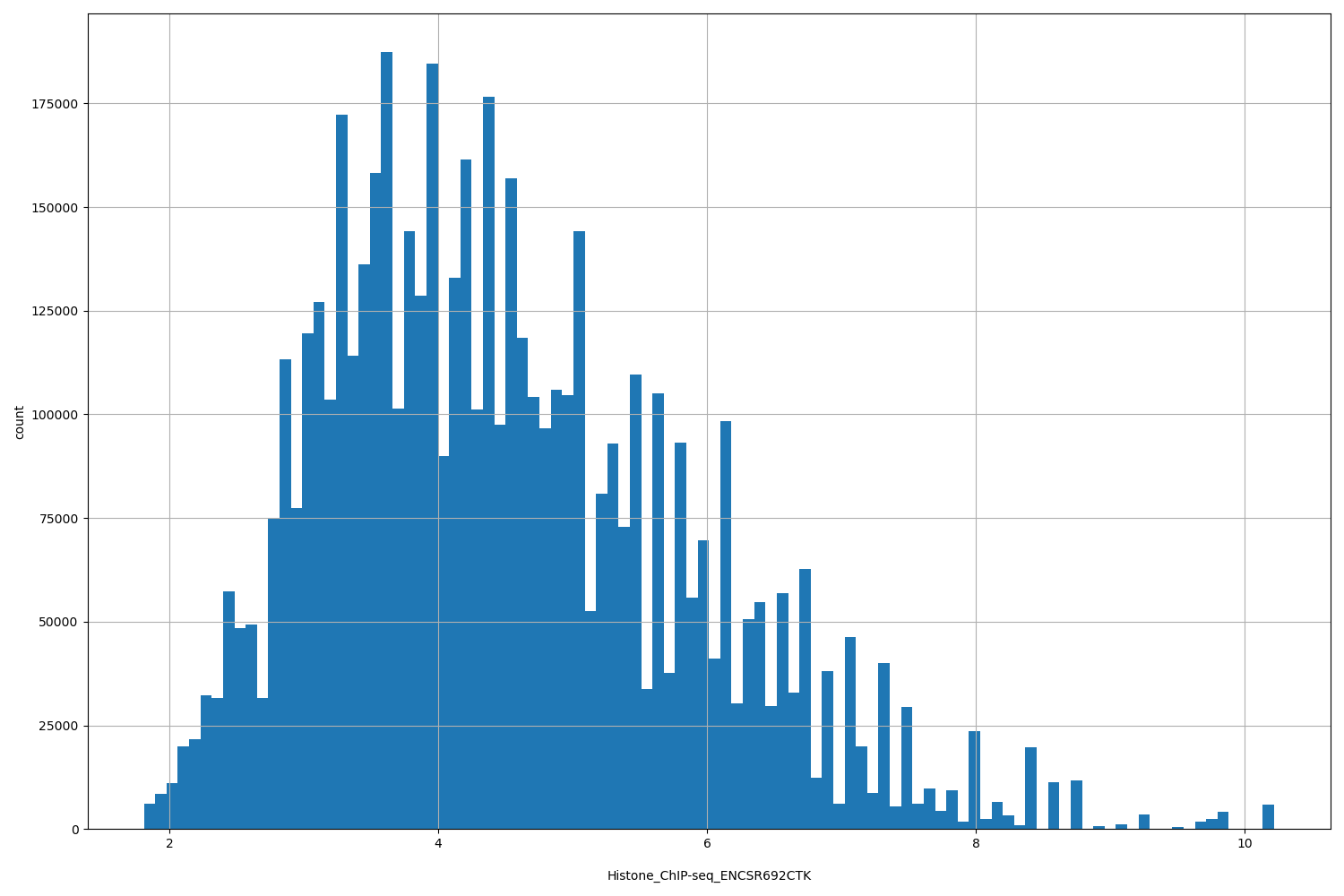
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| Histone_ChIP-seq_ENCSR692CTK | float |
Histone_ChIP-seq_ENCSR692CTK |
Histone_ChIP-seq ENCSR692CTK [biosample_summary="Homo sapiens neuronal stem cell originated from H1" and target="H3K27me3"]
|
 |
[1.81, 10.2] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF153BQX.bed.gz | 205.42 KB | 72bb6c84974f1f078f6e0dd96680f106 |
| ENCFF153BQX.bed.gz.dvc | 100.0 B | bbd59f9100cb6d9d856f3598a09148c3 |
| ENCFF153BQX.tabix.bed.gz | 103.49 KB | 3f8164ec5adadfe3bc3fbc6bd4e7fa24 |
| ENCFF153BQX.tabix.bed.gz.dvc | 106.0 B | e24f0a72d2374aae1f2462b5b0fc523d |
| ENCFF153BQX.tabix.bed.gz.tbi | 42.45 KB | b5b3eee3f0b8ecdaae643a89a78ea0e0 |
| ENCFF153BQX.tabix.bed.gz.tbi.dvc | 109.0 B | 65d962fa01ae670eb3f8020eb77d185f |
| genomic_resource.yaml | 2.68 KB | 489ff5af95a5267d19c505478d00cb03 |
| genomic_resource_original.yaml | 2.55 KB | 5fa8f7cab0e9216977e5cbe1549c94ca |
| statistics/ |