Histone_ChIP-seq_ENCSR659RJP
| Id: | Histone_ChIP-seq/ENCSR659RJP |
| Type: | position_score |
| Version: | 0 |
| Summary: |
HistoneChIP-seq ENCSR659RJP [biosamplesummary="Homo sapiens spleen tissue female adult (53 years)" and target="H3K4me1"] |
| Description: |
status: archived biological_replicates: Rep 1 summary: female adult (53 years) output_type: pseudoreplicated peaks audit_internal_action: Archived analysis {ENCAN996KNX|/analyses/ENCAN996KNX/} has in progress subobject quality standard {encode3-histone-chip|/quality-standards/encode3-histone-chip/} audit_warning: PBC1 (PCR Bottlenecking Coefficient 1, M1/M_distinct) is the ratio of the number of genomic locations where exactly one read maps uniquely (M1) to the number of genomic locations where some reads map (M_distinct). A PBC1 value in the range 0 - 0.5 is severe bottlenecking, 0.5 - 0.8 is moderate bottlenecking, 0.8 - 0.9 is mild bottlenecking, and > 0.9 is no bottlenecking. PBC1 value > 0.9 is recommended, but > 0.8 is acceptable. ENCODE processed alignments file {ENCFF229KGO|/files/ENCFF229KGO/} processed by ChIP-seq ENCODE3 GRCh38 pipeline was generated from a library with PBC1 value of 0.87. audit_warning: PBC2 (PCR Bottlenecking Coefficient 2, M1/M2) is the ratio of the number of genomic locations where exactly one read maps uniquely (M1) to the number of genomic locations where two reads map uniquely (M2). A PBC2 value in the range 0 - 1 is severe bottlenecking, 1 - 3 is moderate bottlenecking, 3 - 10 is mild bottlenecking, > 10 is no bottlenecking. PBC2 value > 10 is recommended, but > 3 is acceptable. ENCODE processed alignments file {ENCFF229KGO|/files/ENCFF229KGO/} processed by ChIP-seq ENCODE3 GRCh38 pipeline was generated from a library with PBC2 value of 7.51. |
| Labels: |
|
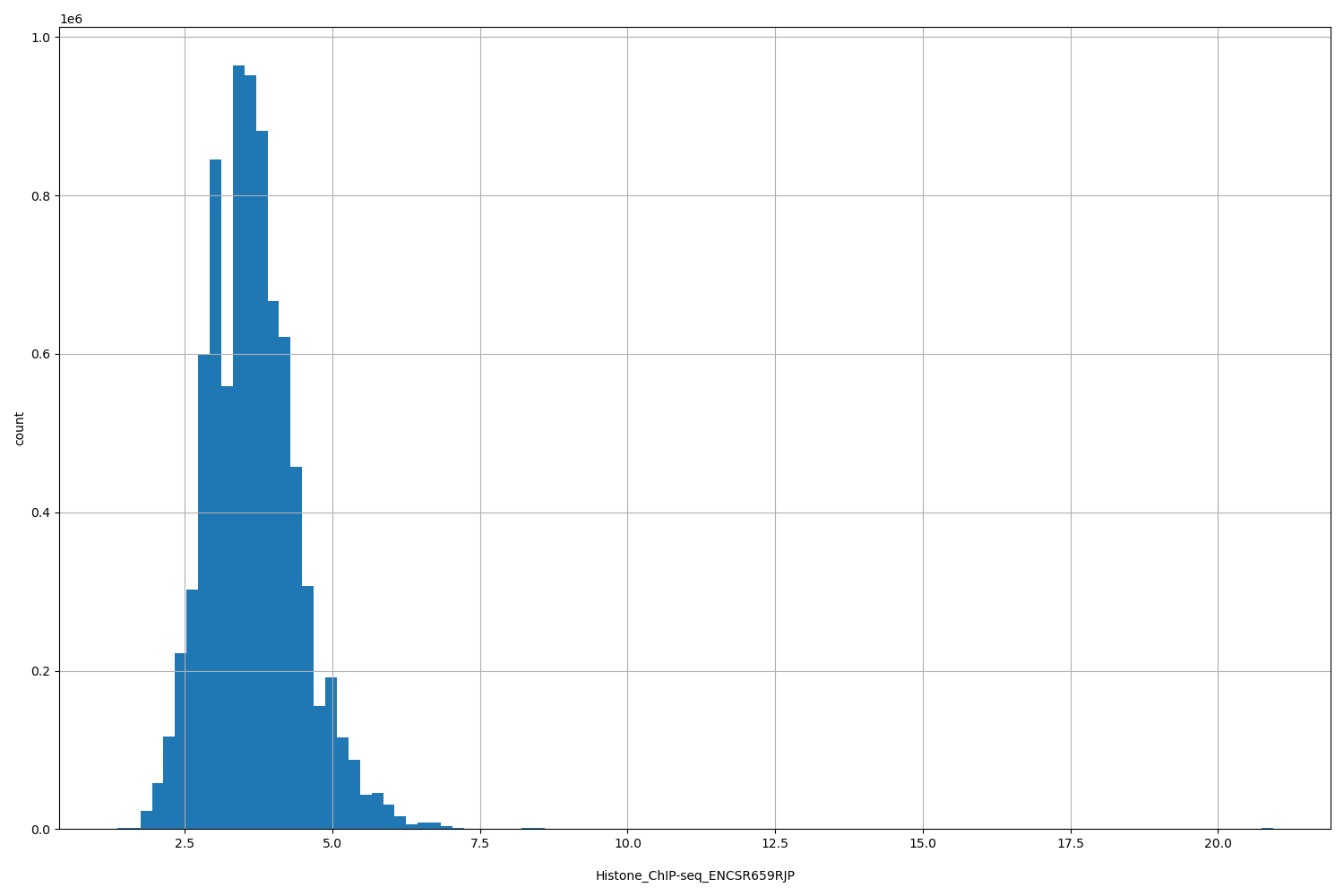
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| Histone_ChIP-seq_ENCSR659RJP | float |
Histone_ChIP-seq_ENCSR659RJP |
Histone_ChIP-seq ENCSR659RJP [biosample_summary="Homo sapiens spleen tissue female adult (53 years)" and target="H3K4me1"]
|
 |
[1.36, 20.9] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF837ZZH.bed.gz | 243.41 KB | 75b622a67ede29d5772ad790d7940e2a |
| ENCFF837ZZH.bed.gz.dvc | 100.0 B | f4a7206cfe0d5195cdb58c26944e8302 |
| ENCFF837ZZH.tabix.bed.gz | 131.45 KB | 6e74058cef2e36e9c5f77a84d7d1d59b |
| ENCFF837ZZH.tabix.bed.gz.dvc | 106.0 B | b08959859d4f1a3b26c926e117eb645f |
| ENCFF837ZZH.tabix.bed.gz.tbi | 85.56 KB | 7d7895b6013bcd5b901a6adb1470ca49 |
| ENCFF837ZZH.tabix.bed.gz.tbi.dvc | 109.0 B | 0fc014f81c8429356b6b23074d33c2fb |
| genomic_resource.yaml | 2.74 KB | aabd070b4f96d71031c19f8e830daec0 |
| genomic_resource_original.yaml | 2.61 KB | 85be348de7ea1fa27e12eb83ae2a0a3c |
| statistics/ |