Histone_ChIP-seq_ENCSR659FAS
| Id: | Histone_ChIP-seq/ENCSR659FAS |
| Type: | position_score |
| Version: | 0 |
| Summary: |
HistoneChIP-seq ENCSR659FAS [biosamplesummary="Homo sapiens chorionic villus tissue female embryo (40 weeks)" and target="H3K4me3"] |
| Description: |
status: released biological_replicates: Rep 1 summary: female embryo (40 weeks) output_type: pseudoreplicated peaks audit_internal_action: Archived analysis {ENCAN559IFK|/analyses/ENCAN559IFK/} has in progress subobject quality standard {encode3-histone-chip|/quality-standards/encode3-histone-chip/} audit_warning: Processed redacted alignments file {ENCFF326OKV|/files/ENCFF326OKV/} processed by ChIP-seq ENCODE3 hg19 pipeline has 16479764 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting H3K4me3-human and investigated as a narrow histone mark is 10 million usable fragments. The recommended value is > 20 million, but > 10 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/}) |
| Labels: |
|
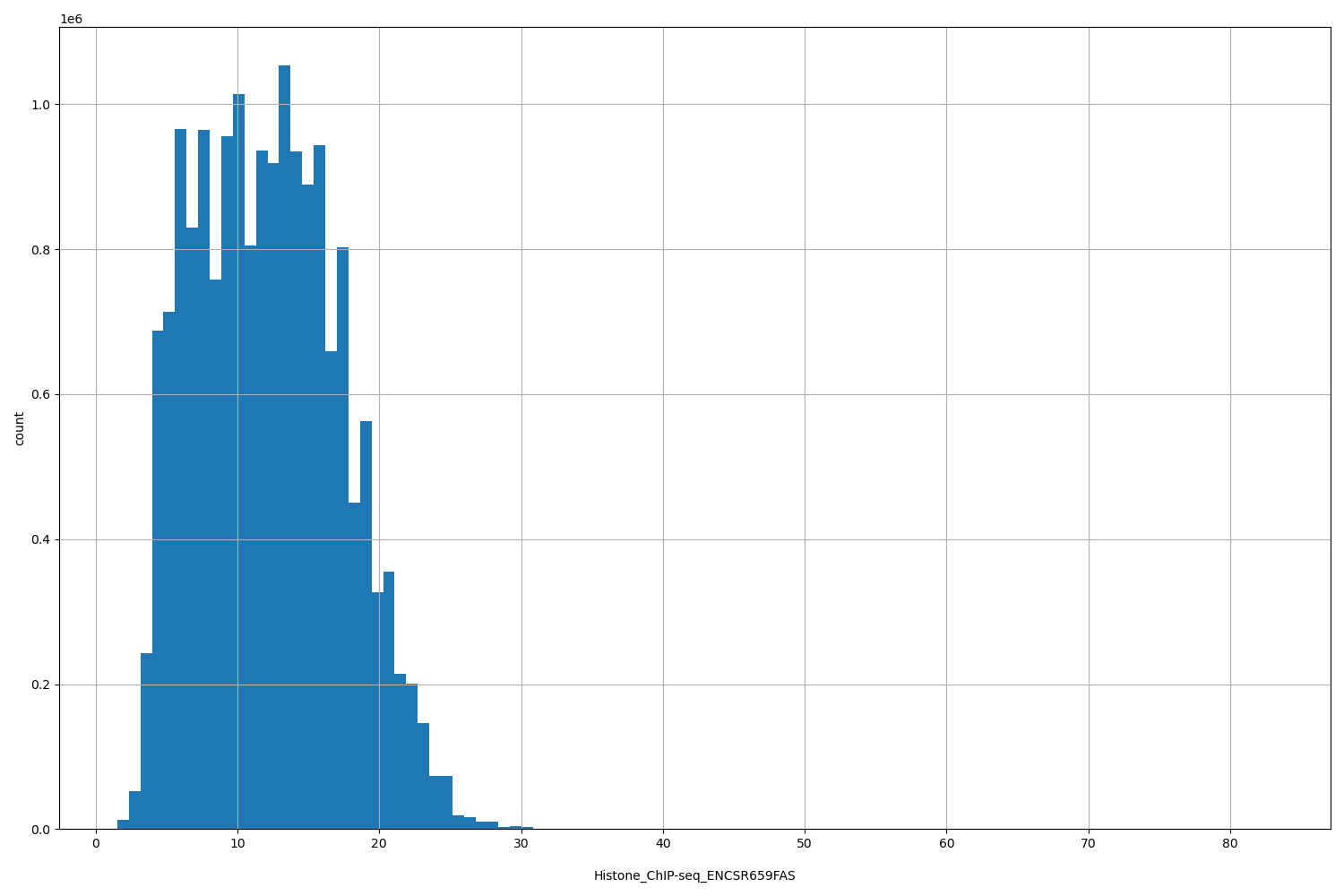
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| Histone_ChIP-seq_ENCSR659FAS | float |
Histone_ChIP-seq_ENCSR659FAS |
Histone_ChIP-seq ENCSR659FAS [biosample_summary="Homo sapiens chorionic villus tissue female embryo (40 weeks)" and target="H3K4me3"]
|
 |
[1.52, 83] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF888FUG.bed.gz | 430.97 KB | f9b8a4ce90dc87ac139b0ac97cb37b62 |
| ENCFF888FUG.bed.gz.dvc | 100.0 B | ef26ad63b31c92de70d047fbfe13f2c0 |
| ENCFF888FUG.tabix.bed.gz | 210.46 KB | 966b1760de92137e3562482954884d8d |
| ENCFF888FUG.tabix.bed.gz.dvc | 106.0 B | 462dc1c7a990c198831b3f205e3f94fd |
| ENCFF888FUG.tabix.bed.gz.tbi | 121.37 KB | ae1a910e9c6d81fede788ad56d38109d |
| ENCFF888FUG.tabix.bed.gz.tbi.dvc | 110.0 B | 1d34d0acefeaa48bd888e262348b3415 |
| genomic_resource.yaml | 2.01 KB | 84385521044de14808c2a53e85cd0674 |
| genomic_resource_original.yaml | 1.87 KB | cda69d6ac29a2c4919c4f76c661e3819 |
| statistics/ |