Histone_ChIP-seq_ENCSR651JDA
| Id: | Histone_ChIP-seq/ENCSR651JDA |
| Type: | position_score |
| Version: | 0 |
| Summary: |
HistoneChIP-seq ENCSR651JDA [biosamplesummary="Homo sapiens smooth muscle cell originated from H9" and target="H3K36me3"] |
| Description: |
status: released biological_replicates: Rep 1, Rep 2 summary: female embryo (5 days) output_type: replicated peaks audit_internal_action: Archived analysis {ENCAN619UVA|/analyses/ENCAN619UVA/} has in progress subobject quality standard {encode3-histone-chip|/quality-standards/encode3-histone-chip/} audit_warning: Processed alignments file {ENCFF343TOI|/files/ENCFF343TOI/} processed by ChIP-seq ENCODE3 GRCh38 pipeline has 39724091 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting H3K36me3-human and investigated as a broad histone mark is 20 million usable fragments. The recommended value is > 45 million, but > 35 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/}) |
| Labels: |
|
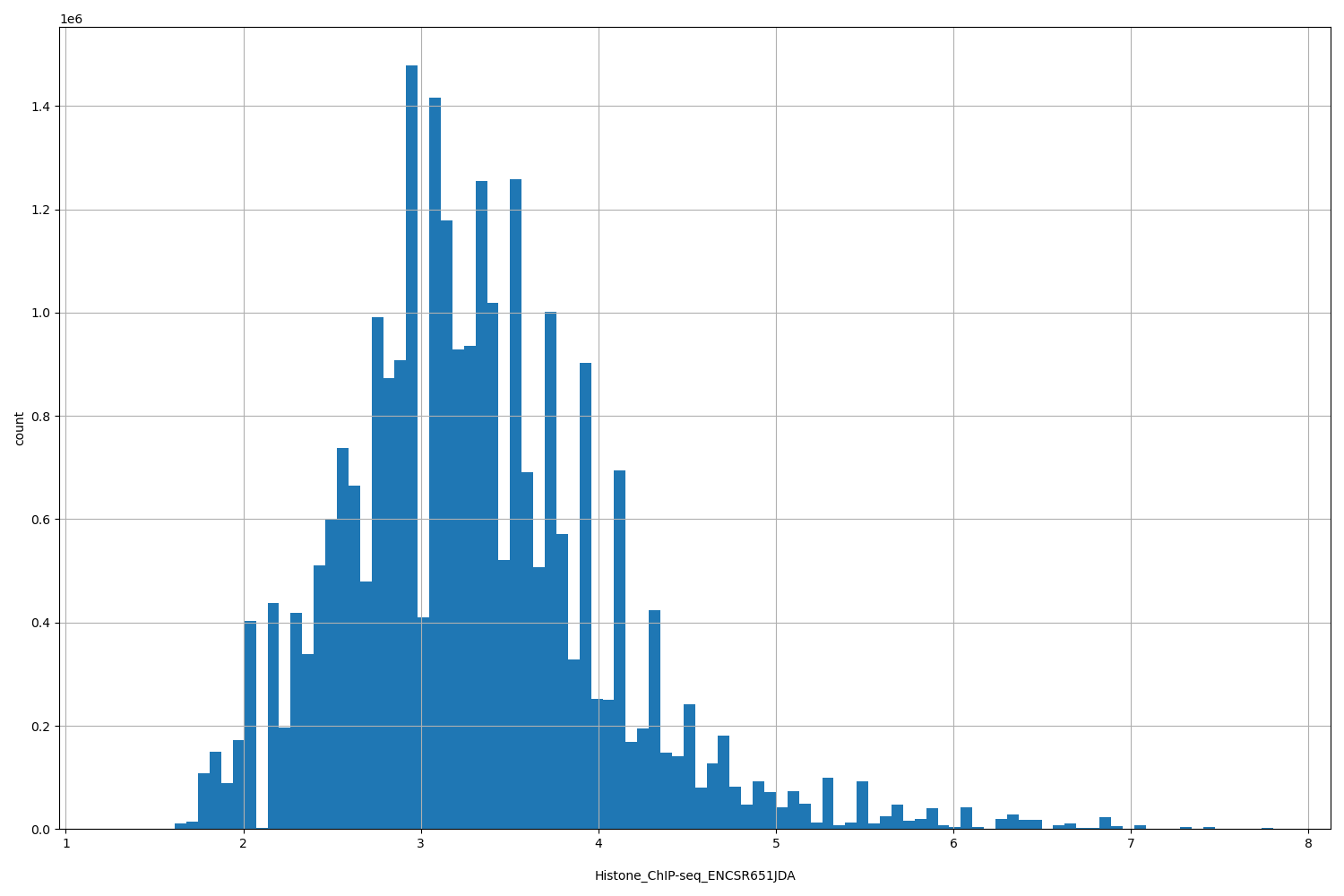
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| Histone_ChIP-seq_ENCSR651JDA | float |
Histone_ChIP-seq_ENCSR651JDA |
Histone_ChIP-seq ENCSR651JDA [biosample_summary="Homo sapiens smooth muscle cell originated from H9" and target="H3K36me3"]
|
 |
[1.29, 7.8] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF823OKF.bed.gz | 1.16 MB | 04598cec389e6d9806999e308c449915 |
| ENCFF823OKF.bed.gz.dvc | 101.0 B | fa08fe6fbf2b86822a0f1a081bb98ea8 |
| ENCFF823OKF.tabix.bed.gz | 600.0 KB | 63565fd36cc17026dfc9461c1c9b782d |
| ENCFF823OKF.tabix.bed.gz.dvc | 106.0 B | 2cfbdb61fee65bf4152c6e203ad38632 |
| ENCFF823OKF.tabix.bed.gz.tbi | 167.81 KB | 2f26bc553d8fa4eb7506785ae198dcec |
| ENCFF823OKF.tabix.bed.gz.tbi.dvc | 110.0 B | 79926f94a20c9fa9c42837d2231d209f |
| genomic_resource.yaml | 1.99 KB | 641dfeeb6f58482c5e701d5e6be9865c |
| genomic_resource_original.yaml | 1.85 KB | 86cff320084edd286552e6cbdb17c604 |
| statistics/ |