Histone_ChIP-seq_ENCSR605SXJ
| Id: | Histone_ChIP-seq/ENCSR605SXJ |
| Type: | position_score |
| Version: | 0 |
| Summary: |
HistoneChIP-seq ENCSR605SXJ [biosamplesummary="Homo sapiens naive thymus-derived CD4-positive, alpha-beta T cell" and target="H3K36me3"] |
| Description: |
status: released biological_replicates: Rep 1 summary: output_type: pseudoreplicated peaks audit_internal_action: Archived analysis {ENCAN787HHE|/analyses/ENCAN787HHE/} has in progress subobject quality standard {encode3-histone-chip|/quality-standards/encode3-histone-chip/} audit_not_compliant: Processed redacted alignments file {ENCFF324OZH|/files/ENCFF324OZH/} processed by ChIP-seq ENCODE3 GRCh38 pipeline has 17984657 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting H3K36me3-human and investigated as a broad histone mark is 20 million usable fragments. The recommended value is > 45 million, but > 35 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/}) audit_warning: PBC1 (PCR Bottlenecking Coefficient 1, M1/M_distinct) is the ratio of the number of genomic locations where exactly one read maps uniquely (M1) to the number of genomic locations where some reads map (M_distinct). A PBC1 value in the range 0 - 0.5 is severe bottlenecking, 0.5 - 0.8 is moderate bottlenecking, 0.8 - 0.9 is mild bottlenecking, and > 0.9 is no bottlenecking. PBC1 value > 0.9 is recommended, but > 0.8 is acceptable. ENCODE processed redacted alignments file {ENCFF324OZH|/files/ENCFF324OZH/} processed by ChIP-seq ENCODE3 GRCh38 pipeline was generated from a library with PBC1 value of 0.90. audit_warning: PBC2 (PCR Bottlenecking Coefficient 2, M1/M2) is the ratio of the number of genomic locations where exactly one read maps uniquely (M1) to the number of genomic locations where two reads map uniquely (M2). A PBC2 value in the range 0 - 1 is severe bottlenecking, 1 - 3 is moderate bottlenecking, 3 - 10 is mild bottlenecking, > 10 is no bottlenecking. PBC2 value > 10 is recommended, but > 3 is acceptable. ENCODE processed redacted alignments file {ENCFF324OZH|/files/ENCFF324OZH/} processed by ChIP-seq ENCODE3 GRCh38 pipeline was generated from a library with PBC2 value of 9.77. |
| Labels: |
|
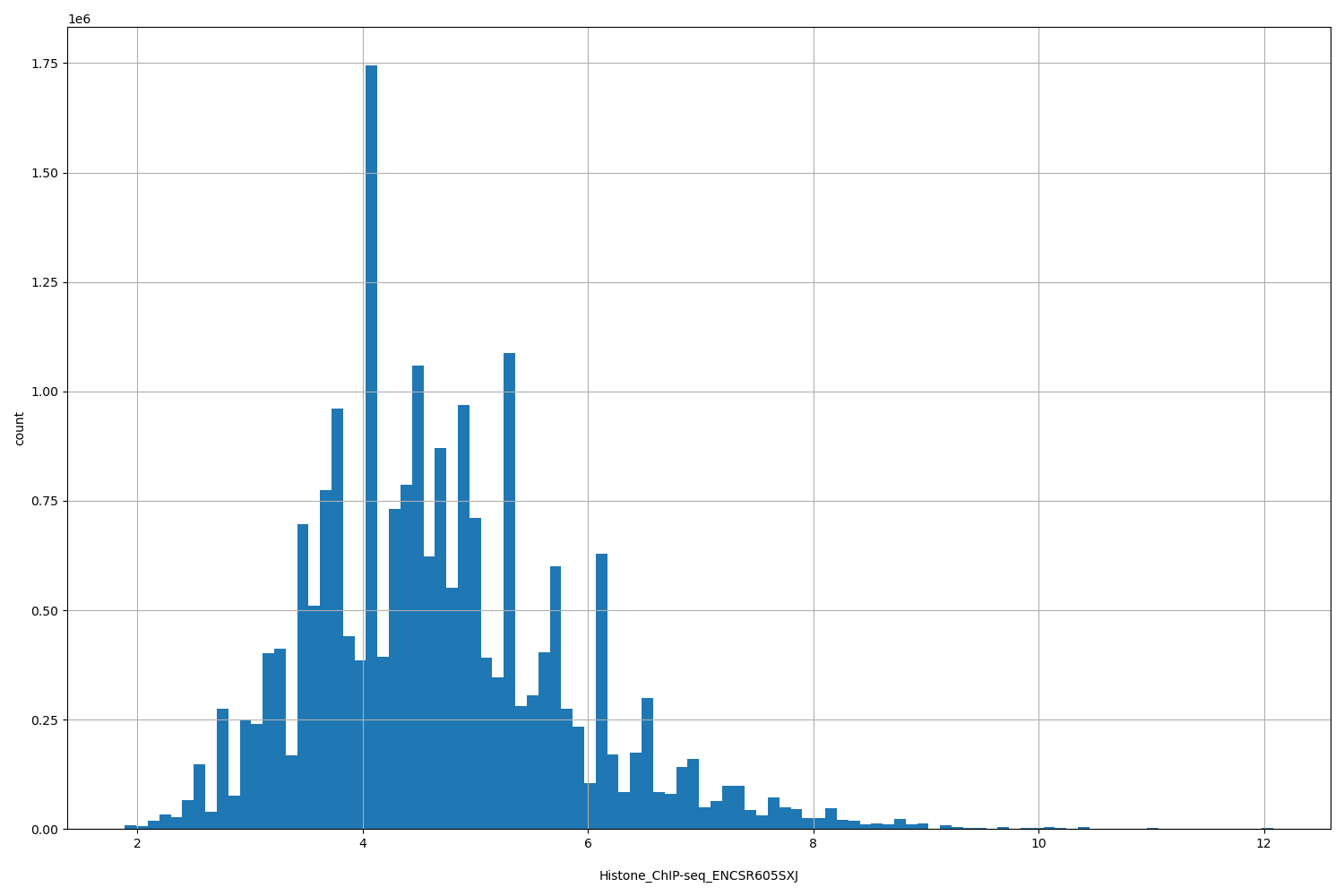
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| Histone_ChIP-seq_ENCSR605SXJ | float |
Histone_ChIP-seq_ENCSR605SXJ |
Histone_ChIP-seq ENCSR605SXJ [biosample_summary="Homo sapiens naive thymus-derived CD4-positive, alpha-beta T cell" and target="H3K36me3"]
|
 |
[1.89, 12.1] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF988ZPL.bed.gz | 680.42 KB | 378db2667baa6af6606623dba644bb17 |
| ENCFF988ZPL.bed.gz.dvc | 100.0 B | 8bc2a6a12be0377696c3b3096c02fd86 |
| ENCFF988ZPL.tabix.bed.gz | 346.44 KB | 288454883e4b1942e7ce52bb3dd264a5 |
| ENCFF988ZPL.tabix.bed.gz.dvc | 106.0 B | 66db340f83474494413fbf018526b0d7 |
| ENCFF988ZPL.tabix.bed.gz.tbi | 113.87 KB | 33aab068e2a79e6a03dbea604b8d4090 |
| ENCFF988ZPL.tabix.bed.gz.tbi.dvc | 110.0 B | 9799060ea82afc0337a271aa0d3d50e1 |
| genomic_resource.yaml | 3.35 KB | fb2b7c6b178e4c826940062ef6702774 |
| genomic_resource_original.yaml | 3.2 KB | 61e41c3fd9bd873deea14a71a2764a42 |
| statistics/ |