Histone_ChIP-seq_ENCSR579TJT
| Id: | Histone_ChIP-seq/ENCSR579TJT |
| Type: | position_score |
| Version: | 0 |
| Summary: |
HistoneChIP-seq ENCSR579TJT [biosamplesummary="Homo sapiens IMR-90" and target="H3K79me1"] |
| Description: |
status: released biological_replicates: Rep 1, Rep 2 summary: output_type: pseudoreplicated peaks audit_internal_action: Released analysis {ENCAN372YYP|/analyses/ENCAN372YYP/} has in progress subobject quality standard {encode4-histone-chip|/quality-standards/encode4-histone-chip/} audit_internal_action: Released analysis {ENCAN372YYP|/analyses/ENCAN372YYP/} has in progress subobject document {1875fd25-6044-4d7d-863e-3fdb18003444|/documents/1875fd25-6044-4d7d-863e-3fdb18003444/} audit_warning: Processed alignments file {ENCFF770OFN|/files/ENCFF770OFN/} processed by ChIP-seq ENCODE4 v1.6.1 GRCh38 pipeline has 13920956 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting H3K79me1-human and investigated as a transcription factor is 10 million usable fragments. The recommended value is > 20 million, but > 10 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/} ) audit_warning: Processed alignments file {ENCFF848ADK|/files/ENCFF848ADK/} processed by ChIP-seq ENCODE4 v1.6.1 GRCh38 pipeline has 13990741 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting H3K79me1-human and investigated as a transcription factor is 10 million usable fragments. The recommended value is > 20 million, but > 10 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/} ) |
| Labels: |
|
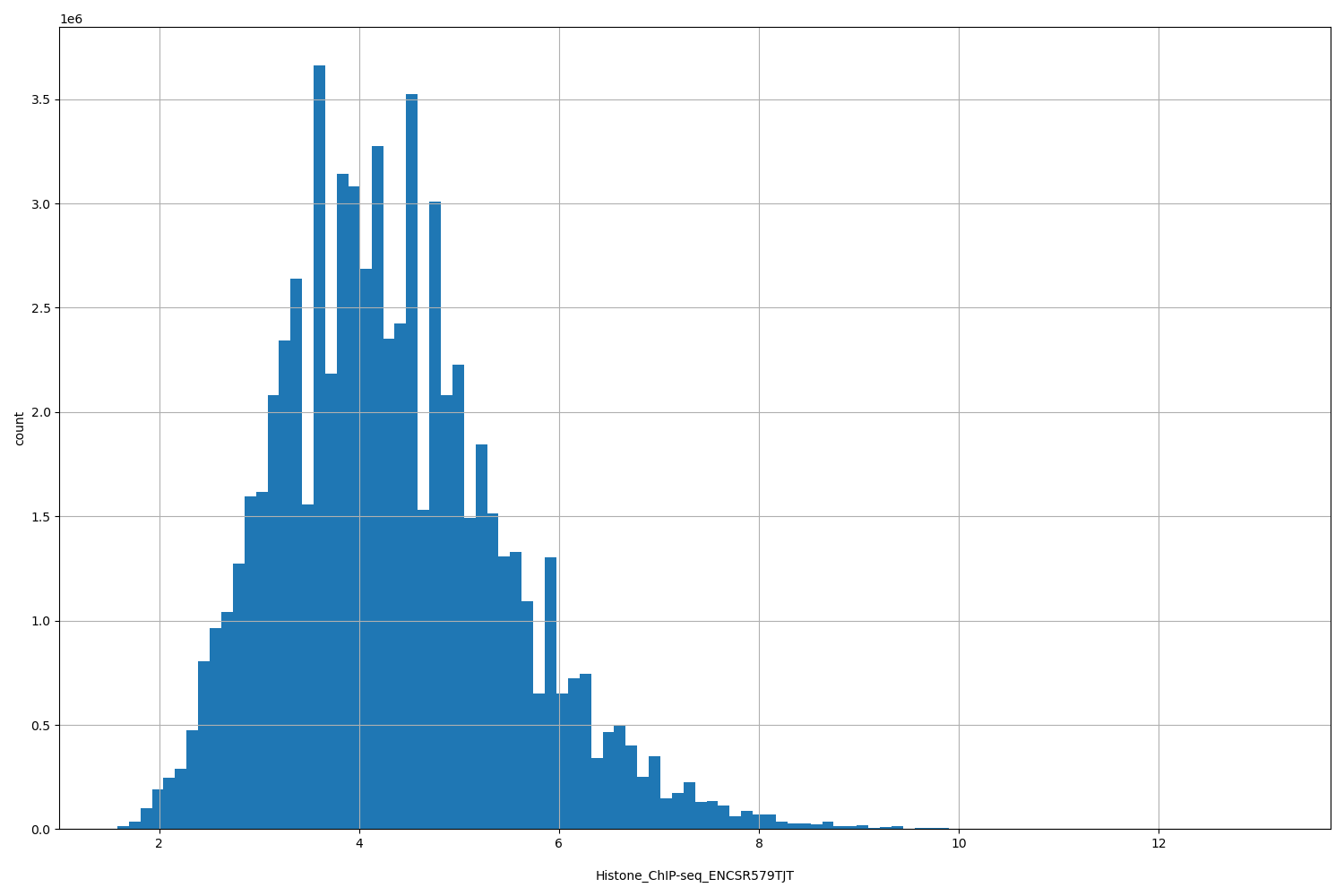
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| Histone_ChIP-seq_ENCSR579TJT | float |
Histone_ChIP-seq_ENCSR579TJT |
Histone_ChIP-seq ENCSR579TJT [biosample_summary="Homo sapiens IMR-90" and target="H3K79me1"]
|
 |
[1.58, 13.1] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF353EDW.bed.gz | 2.85 MB | 80ec52ed83dab61d4f374cc87aa1ea11 |
| ENCFF353EDW.bed.gz.dvc | 101.0 B | ec87ffad6e60c3cd678dec76801e0ebf |
| ENCFF353EDW.tabix.bed.gz | 1.31 MB | f0050bcb778eea56186b179c671b0742 |
| ENCFF353EDW.tabix.bed.gz.dvc | 107.0 B | 275b3b22dcbe4a9e3528fcf691af369c |
| ENCFF353EDW.tabix.bed.gz.tbi | 199.34 KB | 07db118c9423c09cce33b0163d14208f |
| ENCFF353EDW.tabix.bed.gz.tbi.dvc | 110.0 B | 5d94a2c9a6b424b9d3c354e5e037d445 |
| genomic_resource.yaml | 2.6 KB | a232823406fdd6dfb525252232723228 |
| genomic_resource_original.yaml | 2.5 KB | 53e679a0fe530354227861d7572e985c |
| statistics/ |