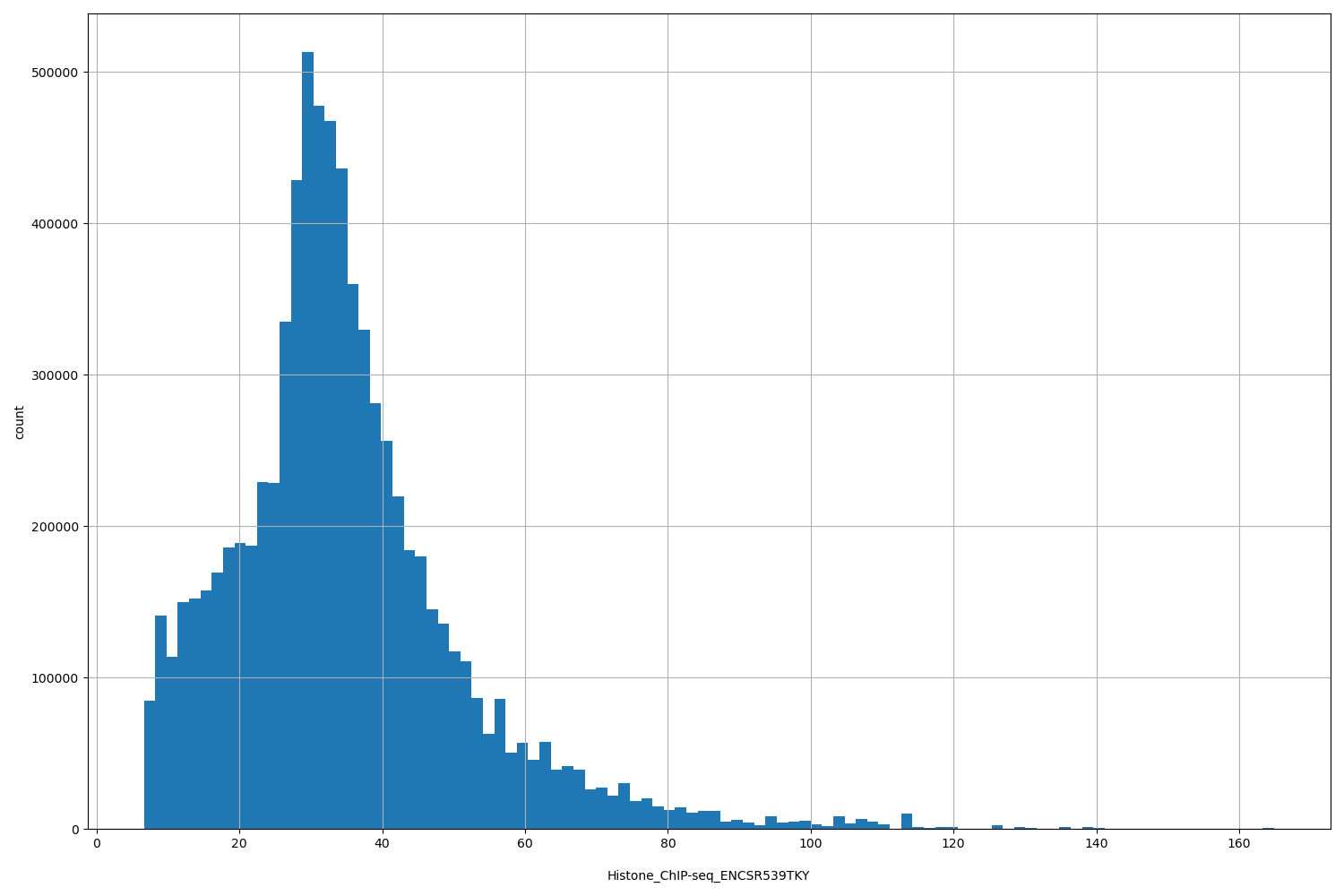
Histone_ChIP-seq_ENCSR539TKY
| Id: | Histone_ChIP-seq/ENCSR539TKY |
| Type: | position_score |
| Version: | 0 |
| Summary: |
HistoneChIP-seq ENCSR539TKY [biosamplesummary="Homo sapiens KMS-11" and target="H3K4me3"] |
| Description: |
status: released biological_replicates: Rep 1, Rep 2 summary: output_type: IDR thresholded peaks audit_internal_action: Released analysis {ENCAN755SSP|/analyses/ENCAN755SSP/} has in progress subobject quality standard {encode4-histone-chip|/quality-standards/encode4-histone-chip/} audit_internal_action: Released analysis {ENCAN755SSP|/analyses/ENCAN755SSP/} has in progress subobject document {1f959b05-4769-47f5-a053-6c02c0a78eee|/documents/1f959b05-4769-47f5-a053-6c02c0a78eee/} audit_not_compliant: Processed alignments file {ENCFF867DKI|/files/ENCFF867DKI/} processed by ChIP-seq ENCODE4 v1.6.1 GRCh38 pipeline has 7597024 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting H3K4me3-human and investigated as a narrow histone mark is 10 million usable fragments. The recommended value is > 20 million, but > 10 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/}) audit_warning: Processed alignments file {ENCFF790PIQ|/files/ENCFF790PIQ/} processed by ChIP-seq ENCODE4 v1.6.1 GRCh38 pipeline has 18110245 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting H3K4me3-human and investigated as a narrow histone mark is 10 million usable fragments. The recommended value is > 20 million, but > 10 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/}) |
| Labels: |
|
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| Histone_ChIP-seq_ENCSR539TKY | float |
Histone_ChIP-seq_ENCSR539TKY |
Histone_ChIP-seq ENCSR539TKY [biosample_summary="Homo sapiens KMS-11" and target="H3K4me3"]
|
 |
[6.62, 165] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF410ORL.bed.gz | 231.12 KB | e2e7f45a0501a1536ba6bffb406851a7 |
| ENCFF410ORL.bed.gz.dvc | 100.0 B | e60c118c53e50db32bf49eb470247ea2 |
| ENCFF410ORL.tabix.bed.gz | 174.24 KB | 29a5e215186398027196ba3fe72982a7 |
| ENCFF410ORL.tabix.bed.gz.dvc | 106.0 B | cd50a04c5fa140c7c1aad358ddedf782 |
| ENCFF410ORL.tabix.bed.gz.tbi | 60.85 KB | e875dc7bea15dd4fa5802e581ec0b37b |
| ENCFF410ORL.tabix.bed.gz.tbi.dvc | 109.0 B | d40a6ca7bbc238a12f984d94e4653627 |
| genomic_resource.yaml | 2.75 KB | 6a9d05980b08a5bbeff0bf506a71a9c2 |
| genomic_resource_original.yaml | 2.64 KB | 5a33d663a852d1c4feaa0d59ccd52ecc |
| statistics/ |