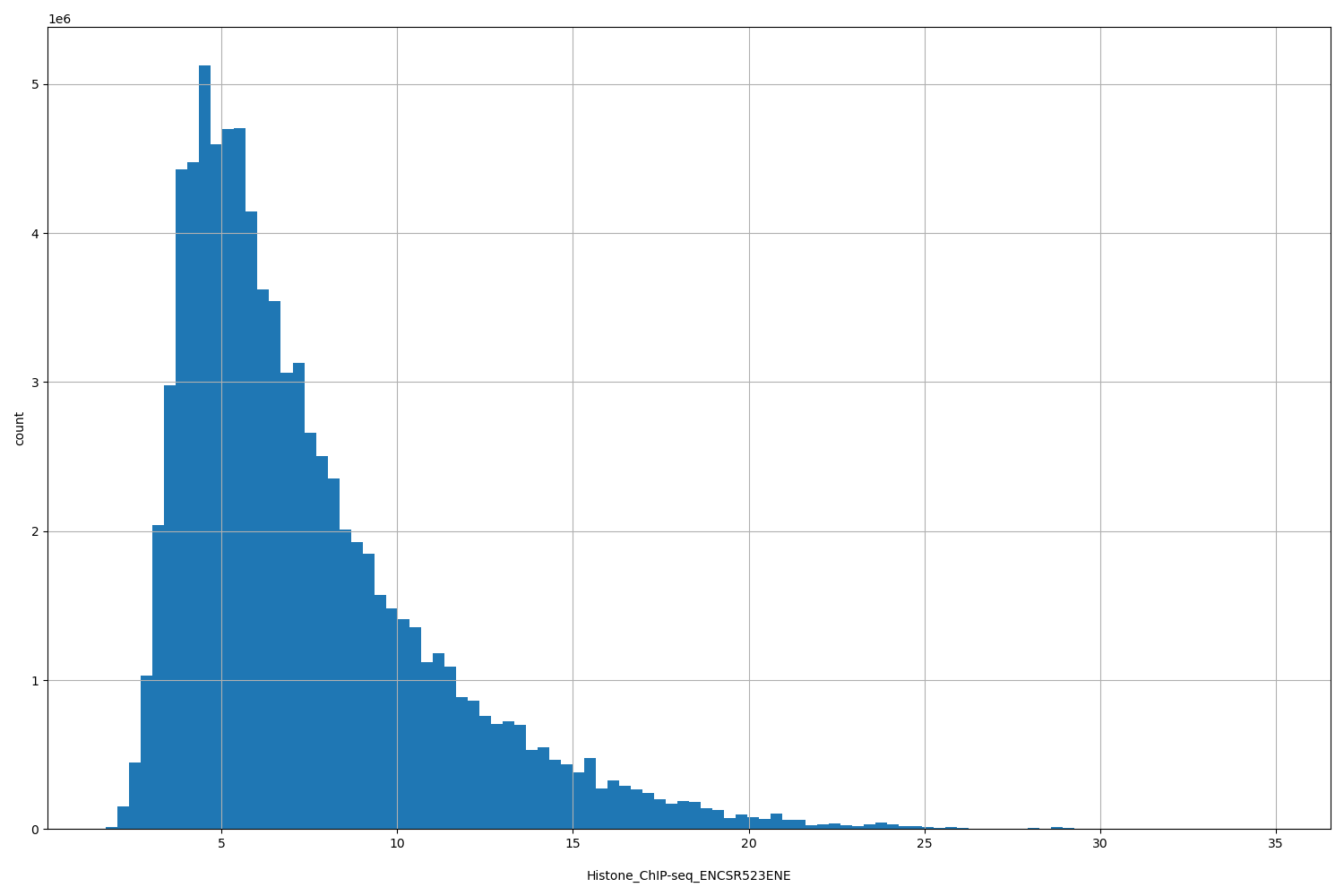
Histone_ChIP-seq_ENCSR523ENE
| Id: | Histone_ChIP-seq/ENCSR523ENE |
| Type: | position_score |
| Version: | 0 |
| Summary: |
HistoneChIP-seq ENCSR523ENE [biosamplesummary="Homo sapiens IMR-90" and target="H2BK12ac"] |
| Description: |
status: released biological_replicates: Rep 1, Rep 2 summary: output_type: pseudoreplicated peaks audit_internal_action: Released analysis {ENCAN390KYZ|/analyses/ENCAN390KYZ/} has in progress subobject document {8a717c3d-f3d9-4010-909c-763d4f4596b2|/documents/8a717c3d-f3d9-4010-909c-763d4f4596b2/} audit_internal_action: Released analysis {ENCAN390KYZ|/analyses/ENCAN390KYZ/} has in progress subobject quality standard {encode4-histone-chip|/quality-standards/encode4-histone-chip/} audit_warning: Processed alignments file {ENCFF053HXY|/files/ENCFF053HXY/} processed by ChIP-seq ENCODE4 v1.6.1 GRCh38 pipeline has 12851123 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting H2BK12ac-human and investigated as a transcription factor is 10 million usable fragments. The recommended value is > 20 million, but > 10 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/} ) audit_warning: Processed alignments file {ENCFF662NJI|/files/ENCFF662NJI/} processed by ChIP-seq ENCODE4 v1.6.1 GRCh38 pipeline has 17445804 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting H2BK12ac-human and investigated as a transcription factor is 10 million usable fragments. The recommended value is > 20 million, but > 10 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/} ) |
| Labels: |
|
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| Histone_ChIP-seq_ENCSR523ENE | float |
Histone_ChIP-seq_ENCSR523ENE |
Histone_ChIP-seq ENCSR523ENE [biosample_summary="Homo sapiens IMR-90" and target="H2BK12ac"]
|
 |
[1.72, 34.9] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF507NUN.bed.gz | 4.44 MB | 2961490dd121333dc6807dc1f29a2db2 |
| ENCFF507NUN.bed.gz.dvc | 101.0 B | e2487066a77252e5a49829075b3e06c4 |
| ENCFF507NUN.tabix.bed.gz | 1.92 MB | d98a442495467816ffc8ff7c73bb7c28 |
| ENCFF507NUN.tabix.bed.gz.dvc | 107.0 B | ca687317f01ef8db63af8068368bc64a |
| ENCFF507NUN.tabix.bed.gz.tbi | 291.12 KB | 961a930e54ae312495977af0471188b3 |
| ENCFF507NUN.tabix.bed.gz.tbi.dvc | 110.0 B | ebb15dcedb216ce3b14a78451466416c |
| genomic_resource.yaml | 2.6 KB | 78ead044dad6103ff938f65d033fa75a |
| genomic_resource_original.yaml | 2.5 KB | d2bf2734b6cc7a6b4a23ef897cd029e4 |
| statistics/ |