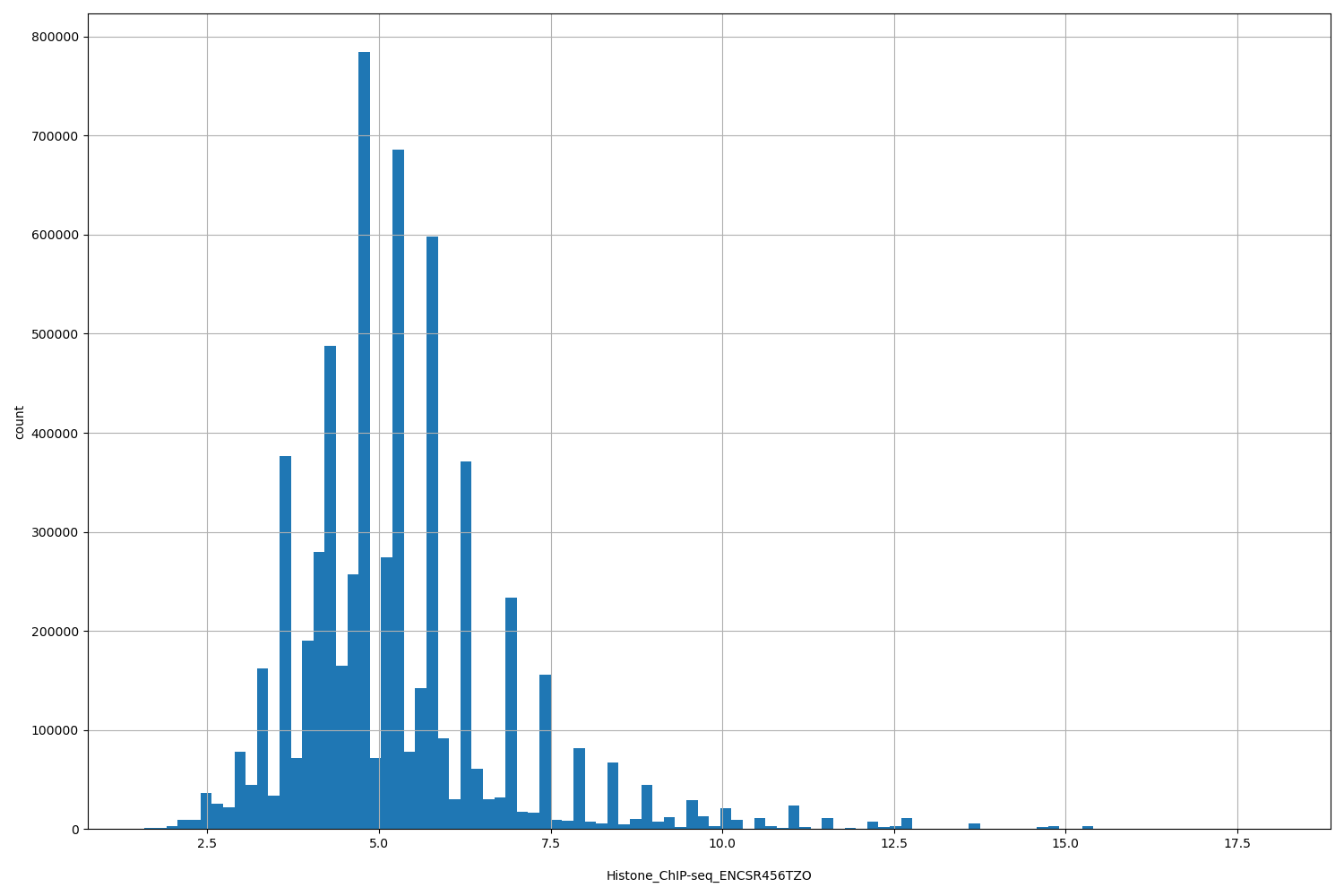
Histone_ChIP-seq_ENCSR456TZO
| Id: | Histone_ChIP-seq/ENCSR456TZO |
| Type: | position_score |
| Version: | 0 |
| Summary: |
HistoneChIP-seq ENCSR456TZO [biosamplesummary="Homo sapiens lung tissue male child (3 years)" and target="H3K9me3"] |
| Description: |
status: released biological_replicates: Rep 1 summary: male child (3 years) output_type: pseudoreplicated peaks audit_internal_action: Archived analysis {ENCAN541GLQ|/analyses/ENCAN541GLQ/} has in progress subobject quality standard {encode3-histone-chip|/quality-standards/encode3-histone-chip/} audit_not_compliant: Processed unfiltered alignments file {ENCFF963ATL|/files/ENCFF963ATL/} processed by ChIP-seq ENCODE3 hg19 pipeline has 28627886 mapped reads. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting H3K9me3-human and investigated as a broad histone mark is 35 million mapped reads. The recommended value is > 45 million, but > 35 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/}) |
| Labels: |
|
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| Histone_ChIP-seq_ENCSR456TZO | float |
Histone_ChIP-seq_ENCSR456TZO |
Histone_ChIP-seq ENCSR456TZO [biosample_summary="Homo sapiens lung tissue male child (3 years)" and target="H3K9me3"]
|
 |
[1.58, 18] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF407SMC.bed.gz | 278.54 KB | 469805a70d03a640f008748105c248b0 |
| ENCFF407SMC.bed.gz.dvc | 100.0 B | a9cc71e61e2025bd976ac65b3c6bc32d |
| ENCFF407SMC.tabix.bed.gz | 145.78 KB | cde0e408887cc2adbb8622c4f30eed86 |
| ENCFF407SMC.tabix.bed.gz.dvc | 106.0 B | b6ecece90f6db2fdd50ce24e3b571d99 |
| ENCFF407SMC.tabix.bed.gz.tbi | 105.07 KB | 1247ab80606a7f9aa060b647985e6eb3 |
| ENCFF407SMC.tabix.bed.gz.tbi.dvc | 110.0 B | 52452d5a44e6c2a71dec4e2ab5dff71e |
| genomic_resource.yaml | 1.96 KB | 155c4261953e7597517c1a41c0af1105 |
| genomic_resource_original.yaml | 1.83 KB | 1eed46808e63117337c67075bb4d4219 |
| statistics/ |