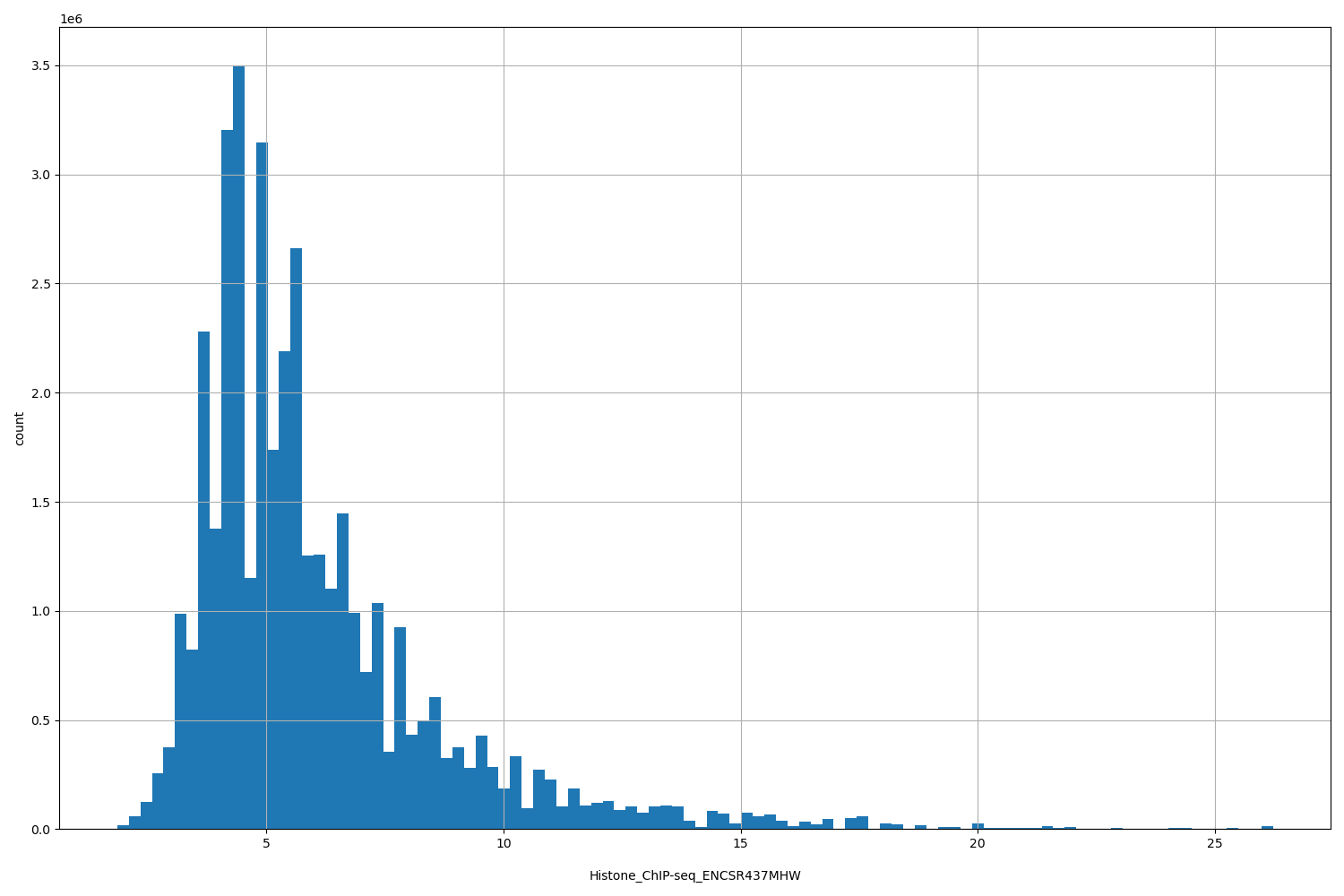
Histone_ChIP-seq_ENCSR437MHW
| Id: | Histone_ChIP-seq/ENCSR437MHW |
| Type: | position_score |
| Version: | 0 |
| Summary: |
HistoneChIP-seq ENCSR437MHW [biosamplesummary="Homo sapiens neutrophil male" and target="H3K9me3"] |
| Description: |
status: released biological_replicates: Rep 1 summary: male output_type: pseudoreplicated peaks audit_internal_action: Archived analysis {ENCAN268SAQ|/analyses/ENCAN268SAQ/} has in progress subobject quality standard {encode3-histone-chip|/quality-standards/encode3-histone-chip/} audit_warning: Processed unfiltered alignments file {ENCFF201RPR|/files/ENCFF201RPR/} processed by ChIP-seq ENCODE3 GRCh38 pipeline has 39071236 mapped reads. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting H3K9me3-human and investigated as a broad histone mark is 35 million mapped reads. The recommended value is > 45 million, but > 35 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/}) |
| Labels: |
|
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| Histone_ChIP-seq_ENCSR437MHW | float |
Histone_ChIP-seq_ENCSR437MHW |
Histone_ChIP-seq ENCSR437MHW [biosample_summary="Homo sapiens neutrophil male" and target="H3K9me3"]
|
 |
[1.86, 26.2] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF523QCG.bed.gz | 1.07 MB | 977b18333bdc792650ef352ea1cb8693 |
| ENCFF523QCG.bed.gz.dvc | 101.0 B | 5fc33bf64f2ebc4484e01b17ffdfb9dd |
| ENCFF523QCG.tabix.bed.gz | 567.16 KB | 620f330662a66651538168b030a87742 |
| ENCFF523QCG.tabix.bed.gz.dvc | 106.0 B | 98a1928691158ee7ccf71ffd976b2904 |
| ENCFF523QCG.tabix.bed.gz.tbi | 249.8 KB | 6c7eb4d9e3ebf7c8efbfec5fc196af8b |
| ENCFF523QCG.tabix.bed.gz.tbi.dvc | 110.0 B | 97ea7c47786f4347cd880247ea7ba7f8 |
| genomic_resource.yaml | 1.87 KB | 06804496223bab3b92c7f4f963529127 |
| genomic_resource_original.yaml | 1.76 KB | 1005da89c1355811ecd5563bf36e5843 |
| statistics/ |