Histone_ChIP-seq_ENCSR431UUY
| Id: | Histone_ChIP-seq/ENCSR431UUY |
| Type: | position_score |
| Version: | 0 |
| Summary: |
HistoneChIP-seq ENCSR431UUY [biosamplesummary="Homo sapiens IMR-90" and target="H3K27me3"] |
| Description: |
status: released biological_replicates: Rep 1, Rep 2 summary: output_type: pseudoreplicated peaks audit_internal_action: Released analysis {ENCAN965OSV|/analyses/ENCAN965OSV/} has in progress subobject quality standard {encode4-histone-chip|/quality-standards/encode4-histone-chip/} audit_internal_action: Released analysis {ENCAN965OSV|/analyses/ENCAN965OSV/} has in progress subobject document {307eb11f-90be-4294-9a77-b570d5755e2a|/documents/307eb11f-90be-4294-9a77-b570d5755e2a/} audit_not_compliant: Processed alignments file {ENCFF547BFL|/files/ENCFF547BFL/} processed by ChIP-seq ENCODE4 v1.6.1 GRCh38 pipeline has 19316653 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting H3K27me3-human and investigated as a broad histone mark is 20 million usable fragments. The recommended value is > 45 million, but > 35 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/}) audit_not_compliant: Processed alignments file {ENCFF084CZB|/files/ENCFF084CZB/} processed by ChIP-seq ENCODE4 v1.6.1 GRCh38 pipeline has 24527220 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting H3K27me3-human and investigated as a broad histone mark is 20 million usable fragments. The recommended value is > 45 million, but > 35 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/}) |
| Labels: |
|
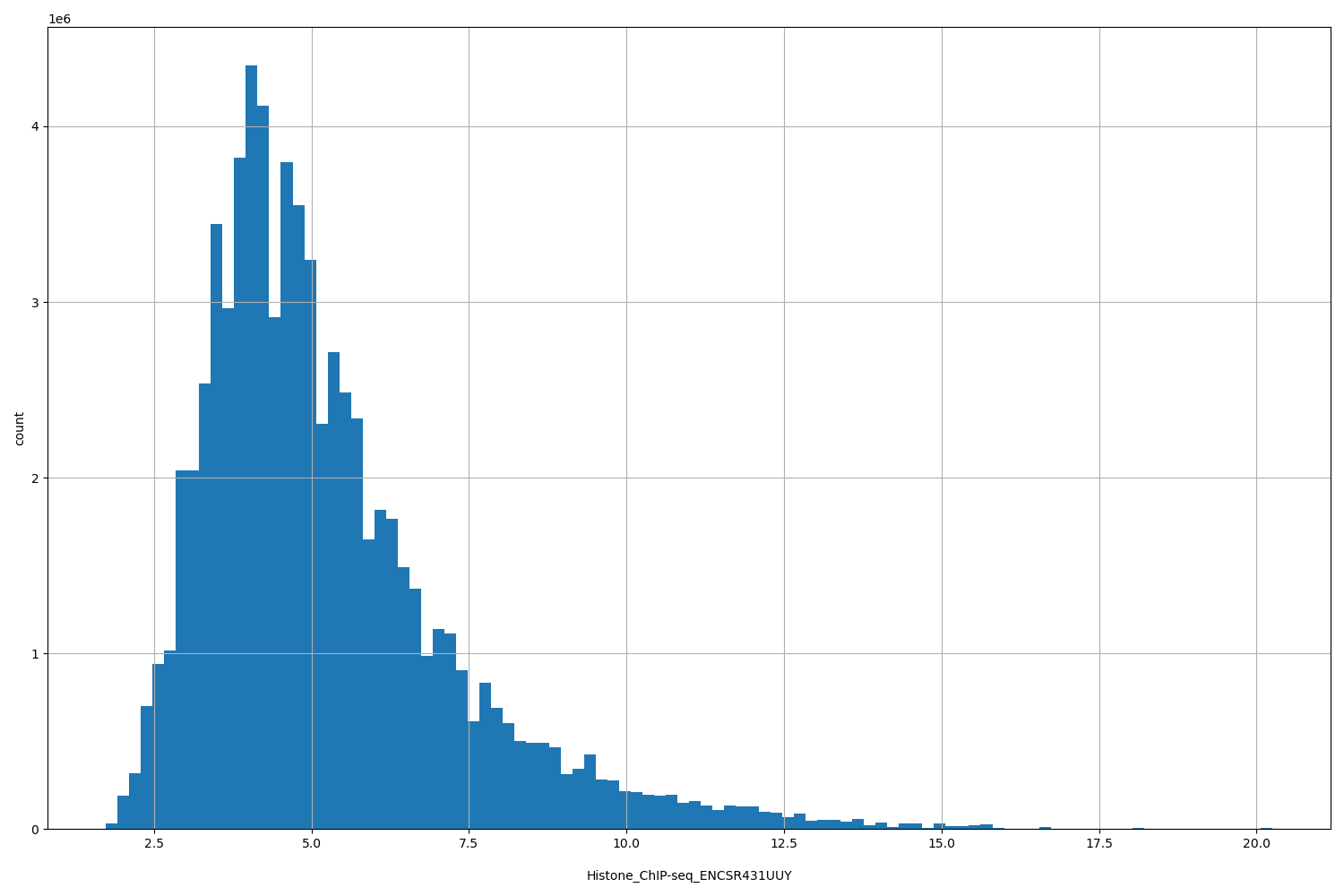
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| Histone_ChIP-seq_ENCSR431UUY | float |
Histone_ChIP-seq_ENCSR431UUY |
Histone_ChIP-seq ENCSR431UUY [biosample_summary="Homo sapiens IMR-90" and target="H3K27me3"]
|
 |
[1.74, 20.2] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF336IXL.bed.gz | 3.54 MB | b5091fa77a952d4ee35759a869c07ac0 |
| ENCFF336IXL.bed.gz.dvc | 101.0 B | 9a3037ccee74e4a9750ab058e9b6c264 |
| ENCFF336IXL.tabix.bed.gz | 1.73 MB | ce8a8a0a386c6be6bf2e2947f54ba02e |
| ENCFF336IXL.tabix.bed.gz.dvc | 107.0 B | 1818d91413e54ee25937ae8c6b92d1c8 |
| ENCFF336IXL.tabix.bed.gz.tbi | 235.62 KB | 206d3082d886c89fdd88acca1fa4fc3a |
| ENCFF336IXL.tabix.bed.gz.tbi.dvc | 110.0 B | 5e7e6822de2ae2007448a826ac9b9e2b |
| genomic_resource.yaml | 2.61 KB | 76b404efaa24fd6a35dc2435fee5804a |
| genomic_resource_original.yaml | 2.5 KB | c27670b4e4a38a158dbac751793ea0e3 |
| statistics/ |