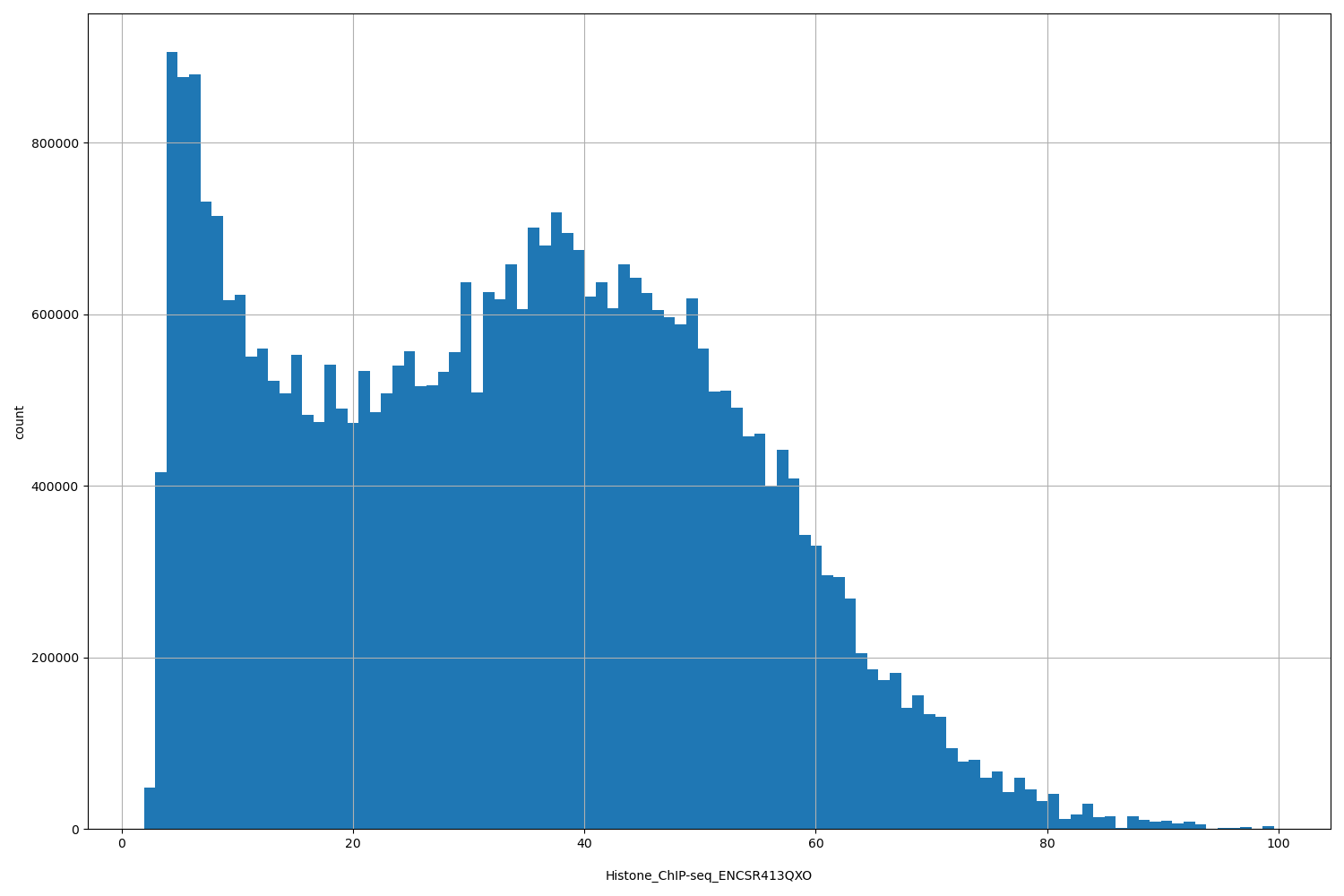
Histone_ChIP-seq_ENCSR413QXO
| Id: | Histone_ChIP-seq/ENCSR413QXO |
| Type: | position_score |
| Version: | 0 |
| Summary: |
HistoneChIP-seq ENCSR413QXO [biosamplesummary="Homo sapiens large intestine tissue male embryo (108 days)" and target="H3K4me3"] |
| Description: |
status: released biological_replicates: Rep 1 summary: male embryo (108 days) output_type: pseudoreplicated peaks audit_internal_action: Archived analysis {ENCAN216QVU|/analyses/ENCAN216QVU/} has in progress subobject quality standard {encode3-histone-chip|/quality-standards/encode3-histone-chip/} audit_warning: Processed alignments file {ENCFF332MLZ|/files/ENCFF332MLZ/} processed by ChIP-seq ENCODE3 GRCh38 pipeline has 19444784 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting H3K4me3-human and investigated as a narrow histone mark is 10 million usable fragments. The recommended value is > 20 million, but > 10 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/}) |
| Labels: |
|
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| Histone_ChIP-seq_ENCSR413QXO | float |
Histone_ChIP-seq_ENCSR413QXO |
Histone_ChIP-seq ENCSR413QXO [biosample_summary="Homo sapiens large intestine tissue male embryo (108 days)" and target="H3K4me3"]
|
 |
[1.91, 99.6] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF190SPX.bed.gz | 825.55 KB | d556c790574ff0a79748d8a0b97e085c |
| ENCFF190SPX.bed.gz.dvc | 100.0 B | 06e6dca3d90937dcb6fd2327397624d2 |
| ENCFF190SPX.tabix.bed.gz | 346.21 KB | a9568be99aa0cdc5189a71a69245580d |
| ENCFF190SPX.tabix.bed.gz.dvc | 106.0 B | 984e9f1c99ea16d60757b95e8db43721 |
| ENCFF190SPX.tabix.bed.gz.tbi | 156.98 KB | 764de5d0e9f4fd8e8dffa3a71e54b6ee |
| ENCFF190SPX.tabix.bed.gz.tbi.dvc | 110.0 B | 18367291e130159a3ac2884d4a4e3cb5 |
| genomic_resource.yaml | 1.99 KB | 8855807513127cd1cbe3d482eaab6dae |
| genomic_resource_original.yaml | 1.85 KB | 929b1302c25c55f5d9571716f9c85c33 |
| statistics/ |