Histone_ChIP-seq_ENCSR366OIQ
| Id: | Histone_ChIP-seq/ENCSR366OIQ |
| Type: | position_score |
| Version: | 0 |
| Summary: |
HistoneChIP-seq ENCSR366OIQ [biosamplesummary="Homo sapiens ectodermal cell originated from HUES64" and target="H3K36me3"] |
| Description: |
status: released biological_replicates: Rep 1 summary: male embryo output_type: pseudoreplicated peaks audit_internal_action: Released analysis {ENCAN301BIZ|/analyses/ENCAN301BIZ/} has in progress subobject quality standard {encode4-histone-chip|/quality-standards/encode4-histone-chip/} audit_not_compliant: Processed redacted alignments file {ENCFF069OJT|/files/ENCFF069OJT/} processed by ChIP-seq ENCODE4 v1.7.0 GRCh38 pipeline has 14090983 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting H3K36me3-human and investigated as a broad histone mark is 20 million usable fragments. The recommended value is > 45 million, but > 35 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/}) |
| Labels: |
|
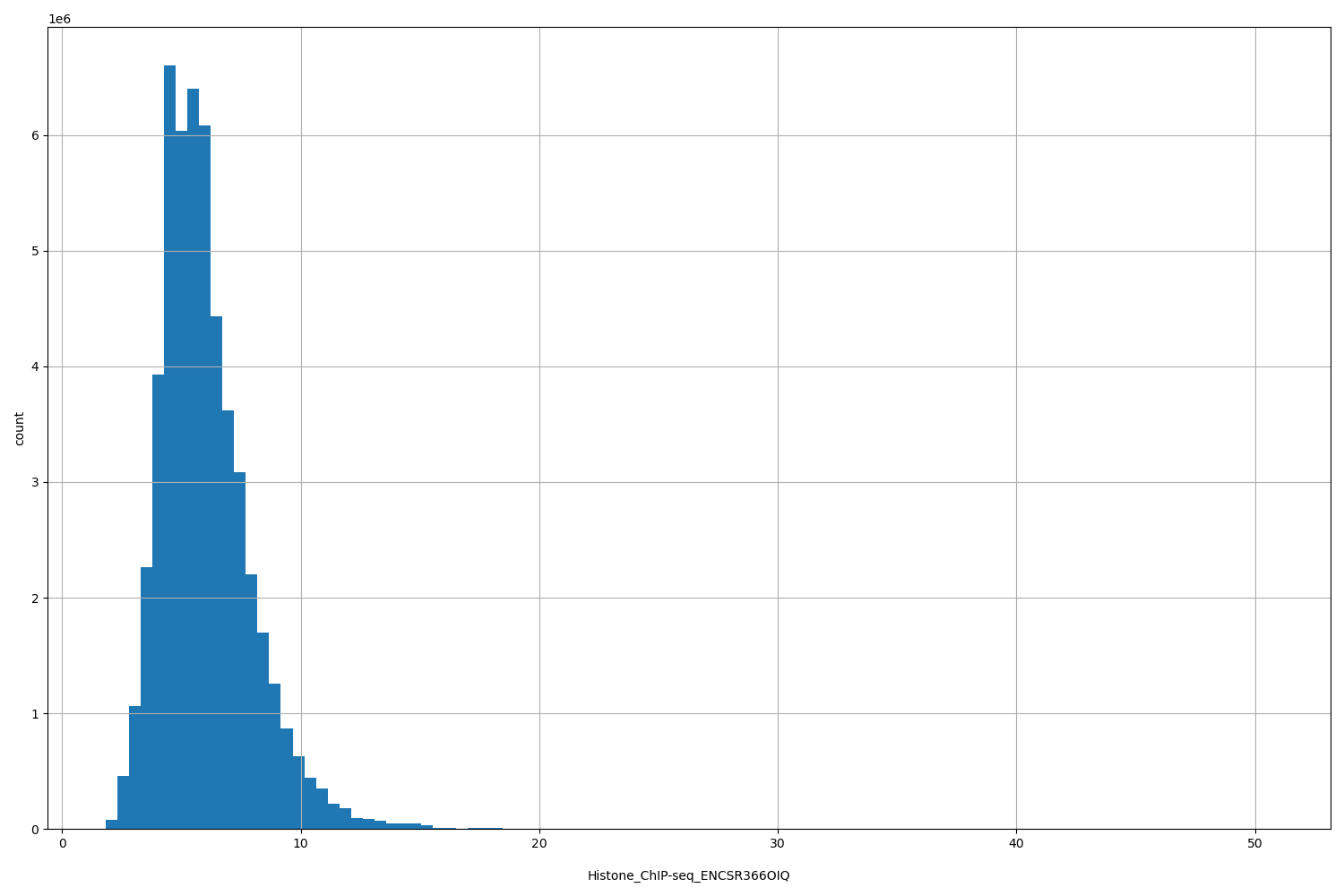
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| Histone_ChIP-seq_ENCSR366OIQ | float |
Histone_ChIP-seq_ENCSR366OIQ |
Histone_ChIP-seq ENCSR366OIQ [biosample_summary="Homo sapiens ectodermal cell originated from HUES64" and target="H3K36me3"]
|
 |
[1.83, 50.7] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF451EGW.bed.gz | 1.21 MB | 6f0a517e1d7eefc9f04a628e49ecbb4e |
| ENCFF451EGW.bed.gz.dvc | 101.0 B | 4d3b5054599ae7ca18fb87f752d850b6 |
| ENCFF451EGW.tabix.bed.gz | 646.56 KB | 1ab048b311a53635a3e57e5fa5beba86 |
| ENCFF451EGW.tabix.bed.gz.dvc | 106.0 B | d6c6214e20653a02a3023dc7327bb855 |
| ENCFF451EGW.tabix.bed.gz.tbi | 144.97 KB | 143deb5cc64f3a862d785579a13d1488 |
| ENCFF451EGW.tabix.bed.gz.tbi.dvc | 110.0 B | 265981b6b4cfc8d21876c643db8736c3 |
| genomic_resource.yaml | 2.0 KB | 90720f32ef6c9a0a7a2f7f2328bf12a1 |
| genomic_resource_original.yaml | 1.86 KB | d94d4a75672874c9ec3b497306c85b65 |
| statistics/ |