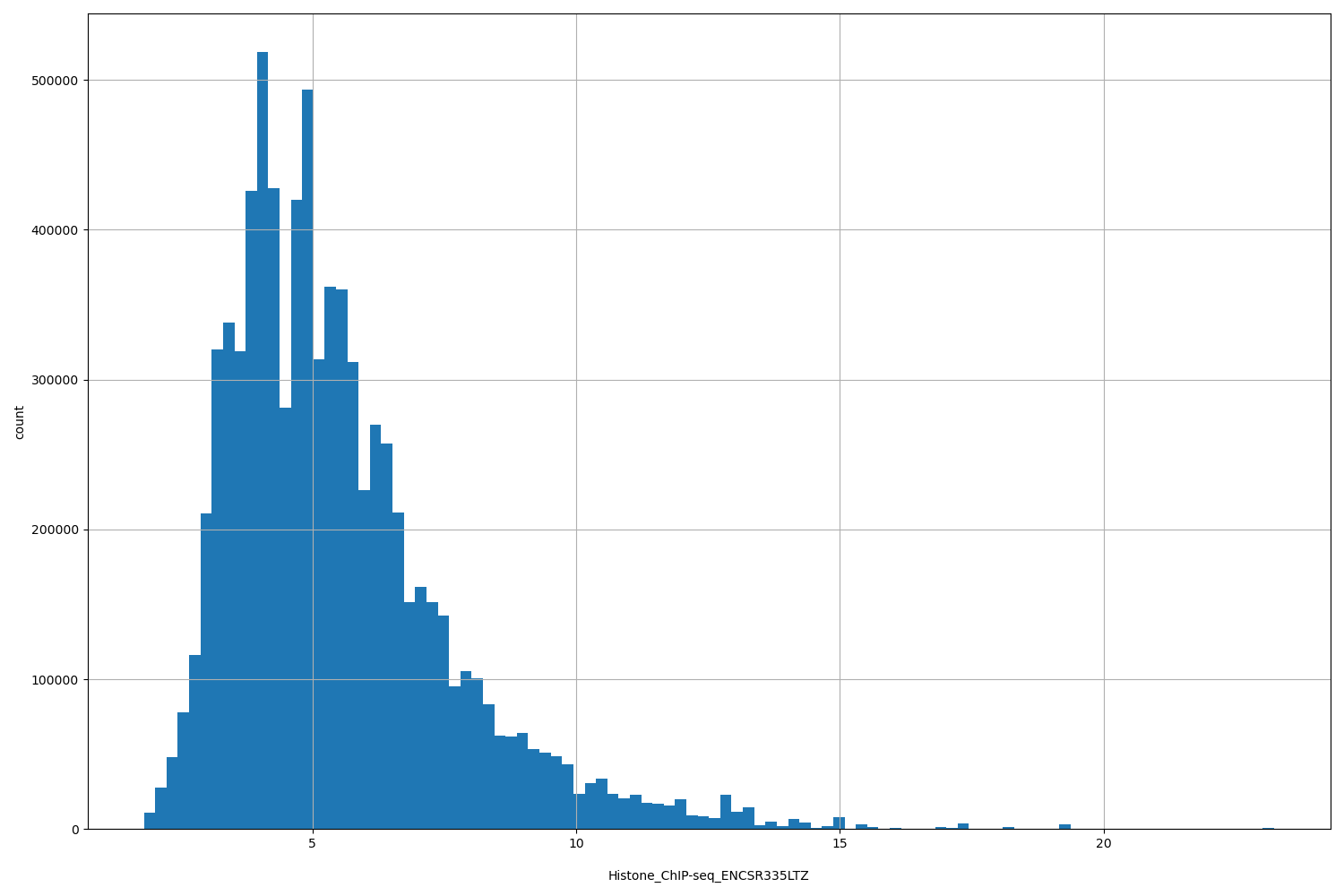
Histone_ChIP-seq_ENCSR335LTZ
| Id: | Histone_ChIP-seq/ENCSR335LTZ |
| Type: | position_score |
| Version: | 0 |
| Summary: |
HistoneChIP-seq ENCSR335LTZ [biosamplesummary="Homo sapiens parathyroid adenoma tissue male adult (65 years)" and target="H3K9ac"] |
| Description: |
status: released biological_replicates: Rep 1 summary: male adult (65 years) output_type: pseudoreplicated peaks audit_internal_action: Archived analysis {ENCAN846PKB|/analyses/ENCAN846PKB/} has in progress subobject quality standard {encode3-histone-chip|/quality-standards/encode3-histone-chip/} audit_warning: PBC1 (PCR Bottlenecking Coefficient 1, M1/M_distinct) is the ratio of the number of genomic locations where exactly one read maps uniquely (M1) to the number of genomic locations where some reads map (M_distinct). A PBC1 value in the range 0 - 0.5 is severe bottlenecking, 0.5 - 0.8 is moderate bottlenecking, 0.8 - 0.9 is mild bottlenecking, and > 0.9 is no bottlenecking. PBC1 value > 0.9 is recommended, but > 0.8 is acceptable. ENCODE processed redacted alignments file {ENCFF902POM|/files/ENCFF902POM/} processed by ChIP-seq ENCODE3 hg19 pipeline was generated from a library with PBC1 value of 0.87. audit_warning: PBC2 (PCR Bottlenecking Coefficient 2, M1/M2) is the ratio of the number of genomic locations where exactly one read maps uniquely (M1) to the number of genomic locations where two reads map uniquely (M2). A PBC2 value in the range 0 - 1 is severe bottlenecking, 1 - 3 is moderate bottlenecking, 3 - 10 is mild bottlenecking, > 10 is no bottlenecking. PBC2 value > 10 is recommended, but > 3 is acceptable. ENCODE processed redacted alignments file {ENCFF902POM|/files/ENCFF902POM/} processed by ChIP-seq ENCODE3 hg19 pipeline was generated from a library with PBC2 value of 7.37. |
| Labels: |
|
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| Histone_ChIP-seq_ENCSR335LTZ | float |
Histone_ChIP-seq_ENCSR335LTZ |
Histone_ChIP-seq ENCSR335LTZ [biosample_summary="Homo sapiens parathyroid adenoma tissue male adult (65 years)" and target="H3K9ac"]
|
 |
[1.81, 23.2] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF012JHM.bed.gz | 268.09 KB | dc9e5894115812c2137b7ea14c1ede1e |
| ENCFF012JHM.bed.gz.dvc | 100.0 B | f7eefc2958a07a6b7ecd43976be1ab41 |
| ENCFF012JHM.tabix.bed.gz | 129.47 KB | e60034ee6e6a9cafcd819b636eb2587e |
| ENCFF012JHM.tabix.bed.gz.dvc | 106.0 B | 9327c5d32a8e7dc2bf8b3708deab3dd1 |
| ENCFF012JHM.tabix.bed.gz.tbi | 85.15 KB | e4df1bf4849004099ca41332de87a42e |
| ENCFF012JHM.tabix.bed.gz.tbi.dvc | 109.0 B | 5cbff4b4a5355a6e0305000245fa6e64 |
| genomic_resource.yaml | 3.3 KB | d439fd5fa977c52250aee73d04eb6131 |
| genomic_resource_original.yaml | 3.16 KB | b89c18afe7958010f532a3a8529ea27c |
| statistics/ |