Histone_ChIP-seq_ENCSR306VSH
| Id: | Histone_ChIP-seq/ENCSR306VSH |
| Type: | position_score |
| Version: | 0 |
| Summary: |
HistoneChIP-seq ENCSR306VSH [biosamplesummary="Homo sapiens Karpas-422" and target="H3K4me1"] |
| Description: |
status: released biological_replicates: Rep 1, Rep 2 summary: output_type: pseudoreplicated peaks audit_internal_action: Released analysis {ENCAN558ERK|/analyses/ENCAN558ERK/} has in progress subobject quality standard {encode4-histone-chip|/quality-standards/encode4-histone-chip/} audit_internal_action: Released analysis {ENCAN558ERK|/analyses/ENCAN558ERK/} has in progress subobject document {b09cc33b-521b-41c6-b011-40379dc7c7bd|/documents/b09cc33b-521b-41c6-b011-40379dc7c7bd/} audit_warning: PBC1 (PCR Bottlenecking Coefficient 1, M1/M_distinct) is the ratio of the number of genomic locations where exactly one read maps uniquely (M1) to the number of genomic locations where some reads map (M_distinct). A PBC1 value in the range 0 - 0.5 is severe bottlenecking, 0.5 - 0.8 is moderate bottlenecking, 0.8 - 0.9 is mild bottlenecking, and > 0.9 is no bottlenecking. PBC1 value > 0.9 is recommended, but > 0.8 is acceptable. ENCODE processed alignments file {ENCFF724PUR|/files/ENCFF724PUR/} processed by ChIP-seq ENCODE4 v1.6.1 GRCh38 pipeline was generated from a library with PBC1 value of 0.85. audit_warning: PBC2 (PCR Bottlenecking Coefficient 2, M1/M2) is the ratio of the number of genomic locations where exactly one read maps uniquely (M1) to the number of genomic locations where two reads map uniquely (M2). A PBC2 value in the range 0 - 1 is severe bottlenecking, 1 - 3 is moderate bottlenecking, 3 - 10 is mild bottlenecking, > 10 is no bottlenecking. PBC2 value > 10 is recommended, but > 3 is acceptable. ENCODE processed alignments file {ENCFF724PUR|/files/ENCFF724PUR/} processed by ChIP-seq ENCODE4 v1.6.1 GRCh38 pipeline was generated from a library with PBC2 value of 6.73. |
| Labels: |
|
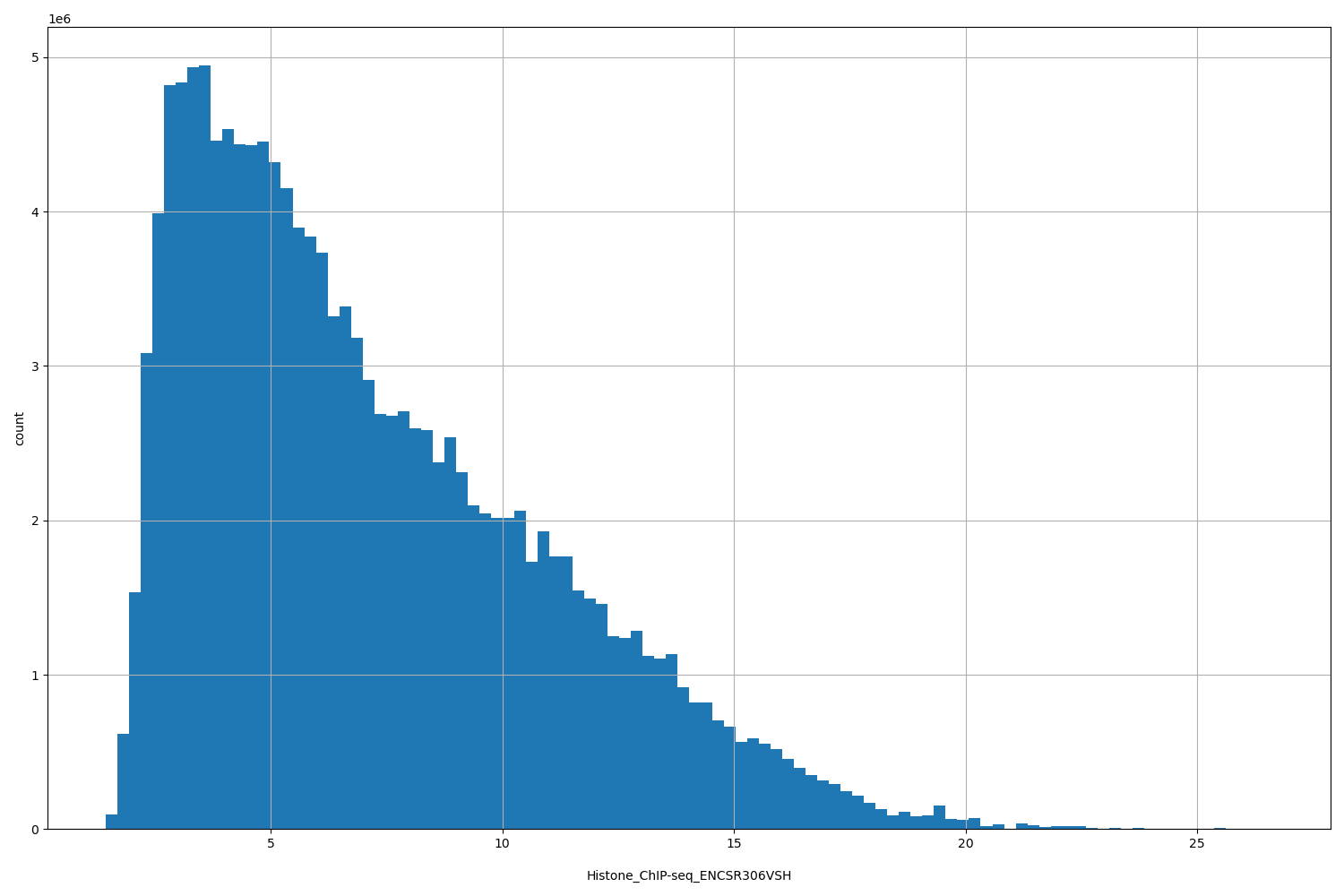
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| Histone_ChIP-seq_ENCSR306VSH | float |
Histone_ChIP-seq_ENCSR306VSH |
Histone_ChIP-seq ENCSR306VSH [biosample_summary="Homo sapiens Karpas-422" and target="H3K4me1"]
|
 |
[1.44, 26.6] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF103BFH.bed.gz | 3.04 MB | 7e0df757b1fbfc7203fc0387b392520e |
| ENCFF103BFH.bed.gz.dvc | 101.0 B | 36cab4605094bcc376ced7f8d360e7a1 |
| ENCFF103BFH.tabix.bed.gz | 1.27 MB | bf7cee022237f94680be444c6aff16b8 |
| ENCFF103BFH.tabix.bed.gz.dvc | 107.0 B | aade6c3173a23aebde99ccad387a32bd |
| ENCFF103BFH.tabix.bed.gz.tbi | 295.97 KB | c950ad65160d0a5c3185297dfde4b7e8 |
| ENCFF103BFH.tabix.bed.gz.tbi.dvc | 110.0 B | cd31a586ff2dda63dcec1ee56b4486e4 |
| genomic_resource.yaml | 2.92 KB | 6583d47e37b3d8b0b6829bfd4c3eff1c |
| genomic_resource_original.yaml | 2.81 KB | b59dc7a22143e645b91310426fe4023c |
| statistics/ |