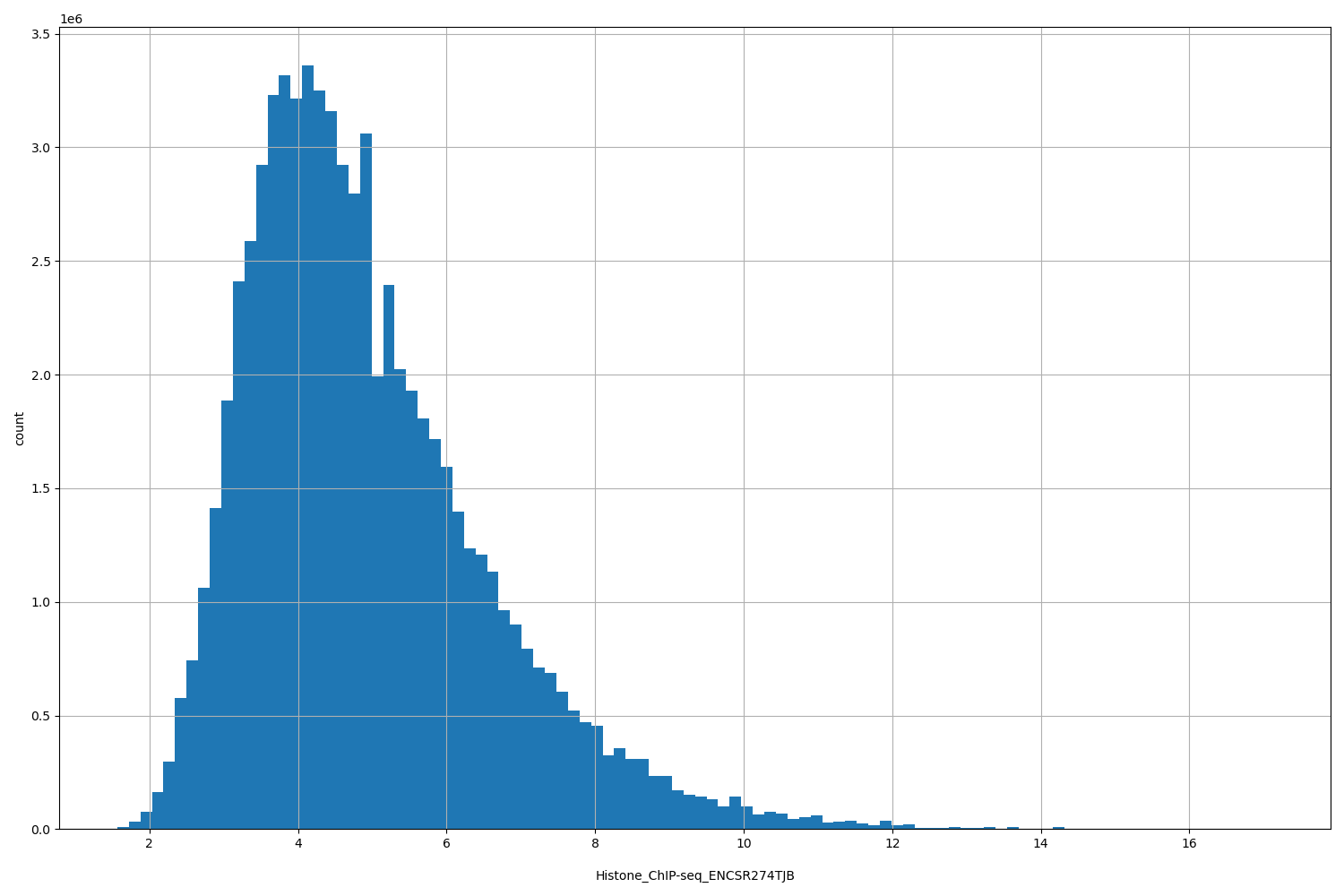
Histone_ChIP-seq_ENCSR274TJB
| Id: | Histone_ChIP-seq/ENCSR274TJB |
| Type: | position_score |
| Version: | 0 |
| Summary: |
HistoneChIP-seq ENCSR274TJB [biosamplesummary="Homo sapiens IMR-90" and target="H2BK15ac"] |
| Description: |
status: released biological_replicates: Rep 1, Rep 2, Rep 3 summary: output_type: pseudoreplicated peaks audit_internal_action: Released analysis {ENCAN718VZG|/analyses/ENCAN718VZG/} has in progress subobject document {f3f80e3a-5877-4295-bf7c-757333271f33|/documents/f3f80e3a-5877-4295-bf7c-757333271f33/} audit_internal_action: Released analysis {ENCAN718VZG|/analyses/ENCAN718VZG/} has in progress subobject quality standard {encode4-histone-chip|/quality-standards/encode4-histone-chip/} audit_warning: Processed alignments file {ENCFF988XSO|/files/ENCFF988XSO/} processed by ChIP-seq ENCODE4 v1.6.1 GRCh38 pipeline has 15928811 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting H2BK15ac-human and investigated as a transcription factor is 10 million usable fragments. The recommended value is > 20 million, but > 10 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/} ) audit_warning: Processed alignments file {ENCFF980RTX|/files/ENCFF980RTX/} processed by ChIP-seq ENCODE4 v1.6.1 GRCh38 pipeline has 11269687 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting H2BK15ac-human and investigated as a transcription factor is 10 million usable fragments. The recommended value is > 20 million, but > 10 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/} ) audit_warning: Processed alignments file {ENCFF510XXG|/files/ENCFF510XXG/} processed by ChIP-seq ENCODE4 v1.6.1 GRCh38 pipeline has 11511089 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting H2BK15ac-human and investigated as a transcription factor is 10 million usable fragments. The recommended value is > 20 million, but > 10 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/} ) |
| Labels: |
|
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| Histone_ChIP-seq_ENCSR274TJB | float |
Histone_ChIP-seq_ENCSR274TJB |
Histone_ChIP-seq ENCSR274TJB [biosample_summary="Homo sapiens IMR-90" and target="H2BK15ac"]
|
 |
[1.57, 17.1] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF840IQF.bed.gz | 3.18 MB | bf80b8252679d6172c58908c2e66879c |
| ENCFF840IQF.bed.gz.dvc | 101.0 B | f9ebf2cae6534ccf4ed2745d8d165efe |
| ENCFF840IQF.tabix.bed.gz | 1.36 MB | 9c425d3e5ffa036603cc6c47fa905be7 |
| ENCFF840IQF.tabix.bed.gz.dvc | 107.0 B | 341e73c656994d279f9955b5677f4cd5 |
| ENCFF840IQF.tabix.bed.gz.tbi | 247.04 KB | 14d54130b59349e6fdbfada2d5ee15db |
| ENCFF840IQF.tabix.bed.gz.tbi.dvc | 110.0 B | 548fb1da74e1740da426b151e00c3407 |
| genomic_resource.yaml | 3.12 KB | 3910c4cd90c64feb4848f71e5ca1050a |
| genomic_resource_original.yaml | 3.01 KB | 4aa8dc8ae72324fccea377d9a368a6a3 |
| statistics/ |