Histone_ChIP-seq_ENCSR269ULZ
| Id: | Histone_ChIP-seq/ENCSR269ULZ |
| Type: | position_score |
| Version: | 0 |
| Summary: |
HistoneChIP-seq ENCSR269ULZ [biosamplesummary="Homo sapiens cardiac muscle cell originated from RUES2" and target="H3K9me3"] |
| Description: |
status: released biological_replicates: Rep 1, Rep 2 summary: embryo output_type: replicated peaks audit_internal_action: Archived analysis {ENCAN801AMI|/analyses/ENCAN801AMI/} has in progress subobject quality standard {encode3-histone-chip|/quality-standards/encode3-histone-chip/} audit_warning: Processed unfiltered alignments file {ENCFF334GUJ|/files/ENCFF334GUJ/} processed by ChIP-seq ENCODE3 hg19 pipeline has 39341160 mapped reads. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting H3K9me3-human and investigated as a broad histone mark is 35 million mapped reads. The recommended value is > 45 million, but > 35 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/}) audit_warning: PBC1 (PCR Bottlenecking Coefficient 1, M1/M_distinct) is the ratio of the number of genomic locations where exactly one read maps uniquely (M1) to the number of genomic locations where some reads map (M_distinct). A PBC1 value in the range 0 - 0.5 is severe bottlenecking, 0.5 - 0.8 is moderate bottlenecking, 0.8 - 0.9 is mild bottlenecking, and > 0.9 is no bottlenecking. PBC1 value > 0.9 is recommended, but > 0.8 is acceptable. ENCODE processed alignments file {ENCFF843MXG|/files/ENCFF843MXG/} processed by ChIP-seq ENCODE3 hg19 pipeline was generated from a library with PBC1 value of 0.86. audit_warning: PBC2 (PCR Bottlenecking Coefficient 2, M1/M2) is the ratio of the number of genomic locations where exactly one read maps uniquely (M1) to the number of genomic locations where two reads map uniquely (M2). A PBC2 value in the range 0 - 1 is severe bottlenecking, 1 - 3 is moderate bottlenecking, 3 - 10 is mild bottlenecking, > 10 is no bottlenecking. PBC2 value > 10 is recommended, but > 3 is acceptable. ENCODE processed alignments file {ENCFF843MXG|/files/ENCFF843MXG/} processed by ChIP-seq ENCODE3 hg19 pipeline was generated from a library with PBC2 value of 7.12. audit_warning: PBC1 (PCR Bottlenecking Coefficient 1, M1/M_distinct) is the ratio of the number of genomic locations where exactly one read maps uniquely (M1) to the number of genomic locations where some reads map (M_distinct). A PBC1 value in the range 0 - 0.5 is severe bottlenecking, 0.5 - 0.8 is moderate bottlenecking, 0.8 - 0.9 is mild bottlenecking, and > 0.9 is no bottlenecking. PBC1 value > 0.9 is recommended, but > 0.8 is acceptable. ENCODE processed alignments file {ENCFF311UHM|/files/ENCFF311UHM/} processed by ChIP-seq ENCODE3 hg19 pipeline was generated from a library with PBC1 value of 0.89. audit_warning: PBC2 (PCR Bottlenecking Coefficient 2, M1/M2) is the ratio of the number of genomic locations where exactly one read maps uniquely (M1) to the number of genomic locations where two reads map uniquely (M2). A PBC2 value in the range 0 - 1 is severe bottlenecking, 1 - 3 is moderate bottlenecking, 3 - 10 is mild bottlenecking, > 10 is no bottlenecking. PBC2 value > 10 is recommended, but > 3 is acceptable. ENCODE processed alignments file {ENCFF311UHM|/files/ENCFF311UHM/} processed by ChIP-seq ENCODE3 hg19 pipeline was generated from a library with PBC2 value of 9.23. |
| Labels: |
|
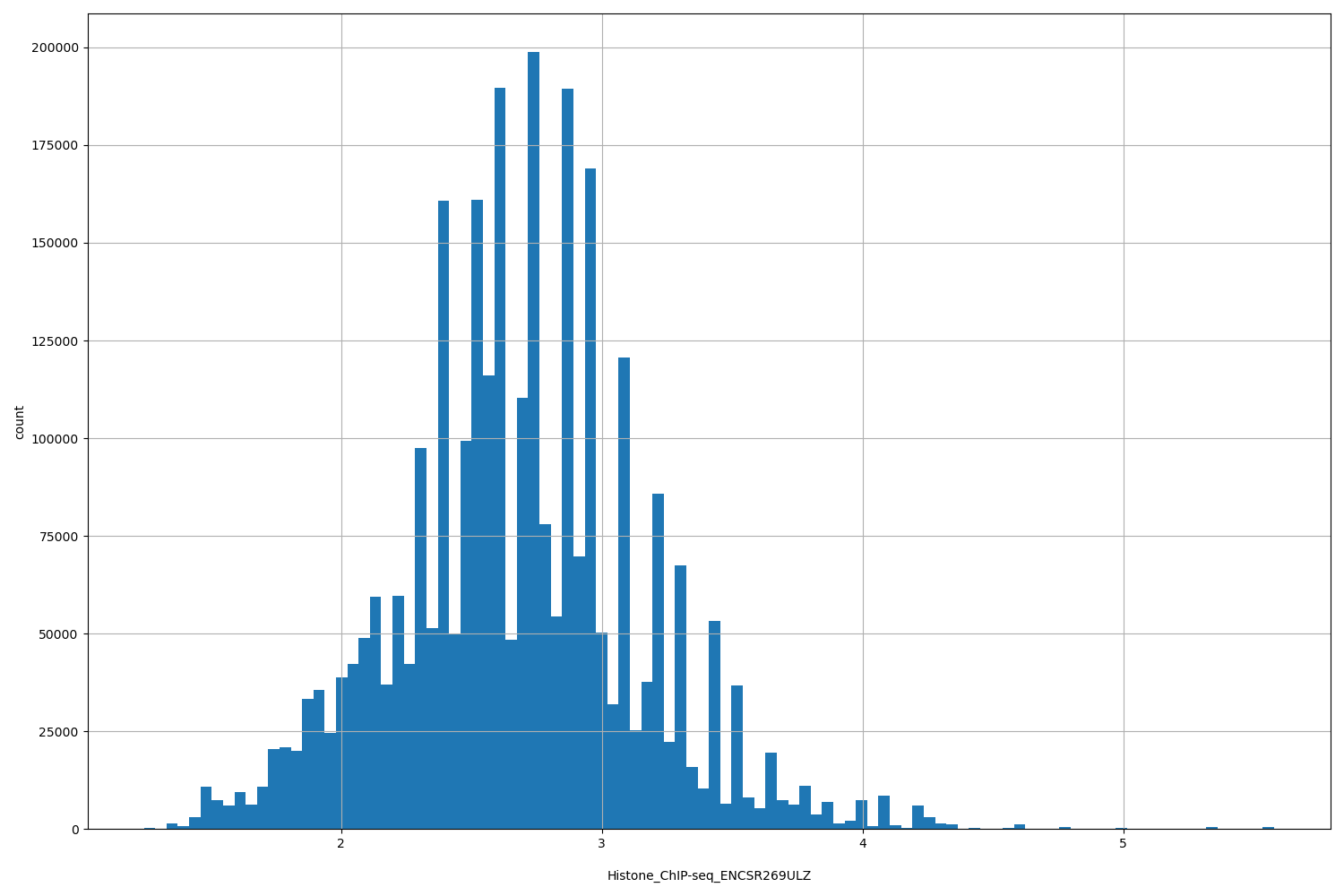
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| Histone_ChIP-seq_ENCSR269ULZ | float |
Histone_ChIP-seq_ENCSR269ULZ |
Histone_ChIP-seq ENCSR269ULZ [biosample_summary="Homo sapiens cardiac muscle cell originated from RUES2" and target="H3K9me3"]
|
 |
[1.24, 5.58] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF817HHD.bed.gz | 140.7 KB | 429b7bd4b859e262c34ab15c38551bc6 |
| ENCFF817HHD.bed.gz.dvc | 100.0 B | a659a111b85aa590cad80f7c3162e8b3 |
| ENCFF817HHD.tabix.bed.gz | 72.54 KB | 13cb67ae349242a87cd0eb46962b0620 |
| ENCFF817HHD.tabix.bed.gz.dvc | 105.0 B | 68287be11a389301688b85fd905bb81f |
| ENCFF817HHD.tabix.bed.gz.tbi | 52.1 KB | ff8ed9e0adfa148a30146507232328e9 |
| ENCFF817HHD.tabix.bed.gz.tbi.dvc | 109.0 B | 5dd90ccdea30026a809a64c5bf04267b |
| genomic_resource.yaml | 4.46 KB | 13988fa16049497422916d5494f3dd76 |
| genomic_resource_original.yaml | 4.33 KB | b793a7a403d84f4994f3a832c8cb2db9 |
| statistics/ |