Histone_ChIP-seq_ENCSR238WMO
| Id: | Histone_ChIP-seq/ENCSR238WMO |
| Type: | position_score |
| Version: | 0 |
| Summary: |
HistoneChIP-seq ENCSR238WMO [biosamplesummary="Homo sapiens neuronal stem cell originated from H9" and target="H3K36me3"] |
| Description: |
status: released biological_replicates: Rep 1, Rep 2 summary: female embryo (5 days) output_type: replicated peaks audit_internal_action: Released analysis {ENCAN075JFE|/analyses/ENCAN075JFE/} has in progress subobject document {8b5ce86d-de7b-4731-af63-389c438bc55e|/documents/8b5ce86d-de7b-4731-af63-389c438bc55e/} audit_internal_action: Released analysis {ENCAN075JFE|/analyses/ENCAN075JFE/} has in progress subobject quality standard {encode4-histone-chip|/quality-standards/encode4-histone-chip/} audit_not_compliant: Processed alignments file {ENCFF508MTC|/files/ENCFF508MTC/} processed by ChIP-seq ENCODE4 v1.6.1 GRCh38 pipeline has 32338538 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting H3K36me3-human and investigated as a broad histone mark is 20 million usable fragments. The recommended value is > 45 million, but > 35 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/}) audit_not_compliant: Processed alignments file {ENCFF246DZP|/files/ENCFF246DZP/} processed by ChIP-seq ENCODE4 v1.6.1 GRCh38 pipeline has 28358516 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting H3K36me3-human and investigated as a broad histone mark is 20 million usable fragments. The recommended value is > 45 million, but > 35 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/}) |
| Labels: |
|
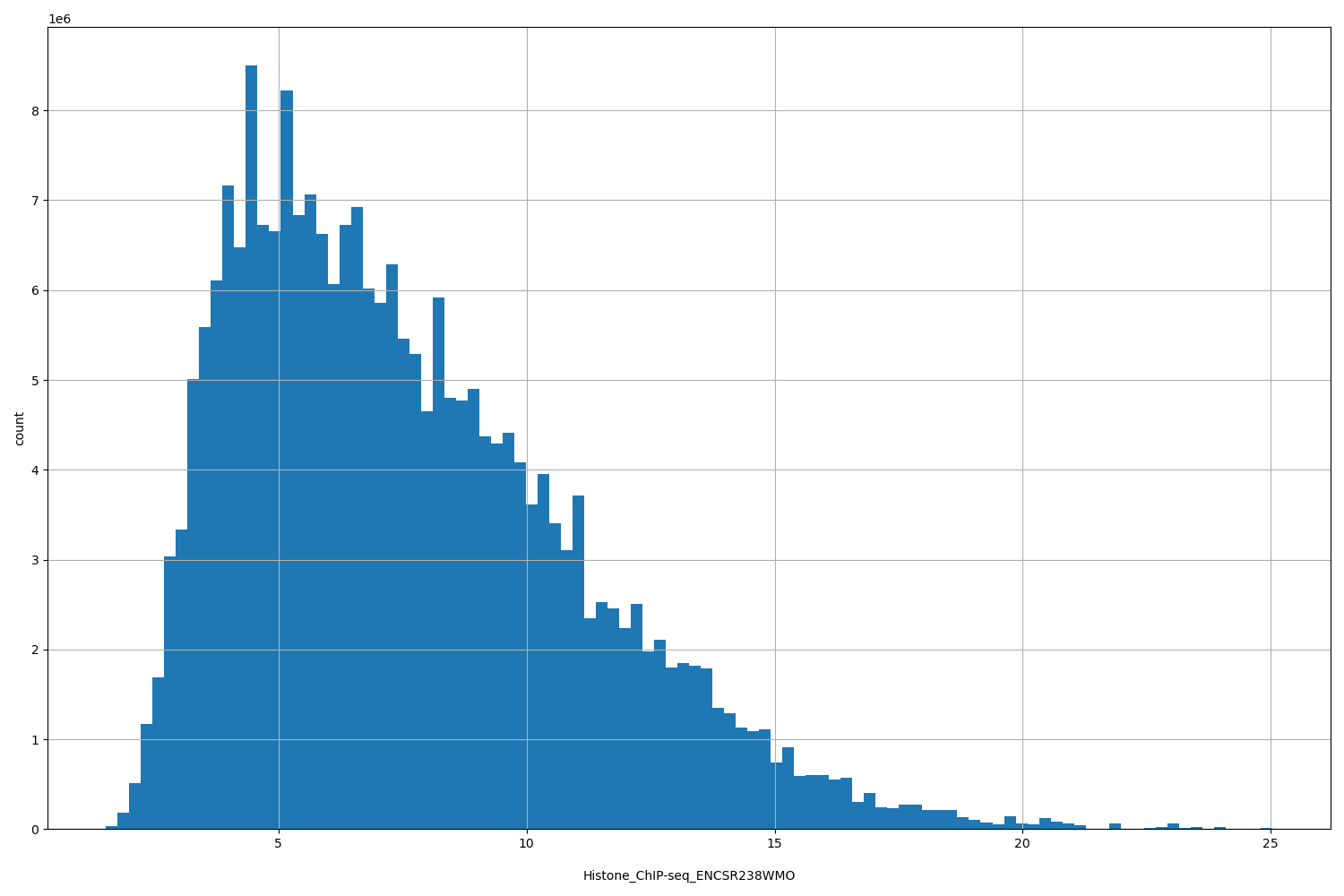
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| Histone_ChIP-seq_ENCSR238WMO | float |
Histone_ChIP-seq_ENCSR238WMO |
Histone_ChIP-seq ENCSR238WMO [biosample_summary="Homo sapiens neuronal stem cell originated from H9" and target="H3K36me3"]
|
 |
[1.52, 25] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF133ROS.bed.gz | 3.62 MB | 352545a6652e092b4e4dcef303ded514 |
| ENCFF133ROS.bed.gz.dvc | 101.0 B | d5b7ebe47544f0a63a6a4bd1ea8c9de1 |
| ENCFF133ROS.tabix.bed.gz | 1.63 MB | 3e774d3f46b0f27c82f0627e3426caa7 |
| ENCFF133ROS.tabix.bed.gz.dvc | 107.0 B | 5e599b48c7c5a4d95621e2a17b4738d0 |
| ENCFF133ROS.tabix.bed.gz.tbi | 156.04 KB | fee44e993ee483e1474b053a0f692014 |
| ENCFF133ROS.tabix.bed.gz.tbi.dvc | 110.0 B | 5cad64aaf9c28676c75c5c2b0020ba24 |
| genomic_resource.yaml | 2.83 KB | f7b96b726298ebe8745c98c4c675bf7f |
| genomic_resource_original.yaml | 2.7 KB | 7b1f5a798a4c4ea709cb986563009c6f |
| statistics/ |