Histone_ChIP-seq_ENCSR231FDF
| Id: | Histone_ChIP-seq/ENCSR231FDF |
| Type: | position_score |
| Version: | 0 |
| Summary: |
HistoneChIP-seq ENCSR231FDF [biosamplesummary="Homo sapiens CD8-positive, alpha-beta T cell" and target="H3K4me3"] |
| Description: |
status: released biological_replicates: Rep 1, Rep 2 summary: output_type: pseudoreplicated peaks audit_internal_action: Released analysis {ENCAN295RWM|/analyses/ENCAN295RWM/} has in progress subobject quality standard {encode4-histone-chip|/quality-standards/encode4-histone-chip/} audit_warning: Processed redacted alignments file {ENCFF736WZO|/files/ENCFF736WZO/} processed by ChIP-seq ENCODE4 v1.7.0 GRCh38 pipeline has 15236093 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting H3K4me3-human and investigated as a narrow histone mark is 10 million usable fragments. The recommended value is > 20 million, but > 10 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/}) |
| Labels: |
|
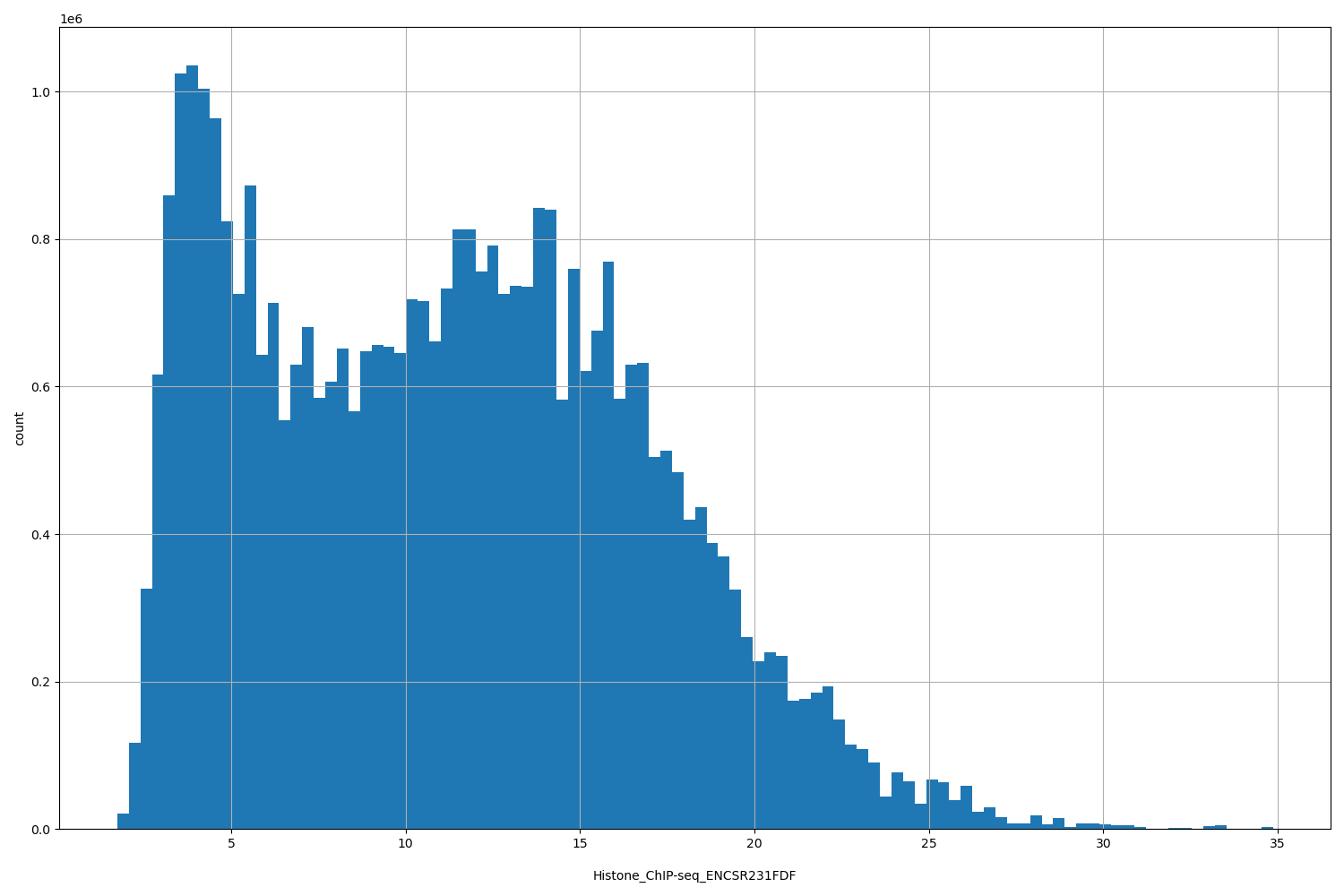
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| Histone_ChIP-seq_ENCSR231FDF | float |
Histone_ChIP-seq_ENCSR231FDF |
Histone_ChIP-seq ENCSR231FDF [biosample_summary="Homo sapiens CD8-positive, alpha-beta T cell" and target="H3K4me3"]
|
 |
[1.74, 34.9] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF785IPD.bed.gz | 814.27 KB | 060e7f570a85110b7d5ed045842d15a5 |
| ENCFF785IPD.bed.gz.dvc | 100.0 B | 913e836ccee0939a2458952e74a732a9 |
| ENCFF785IPD.tabix.bed.gz | 347.66 KB | 4199d4926f331e54634f0c2176f63287 |
| ENCFF785IPD.tabix.bed.gz.dvc | 106.0 B | f1580dad50c60b261c994e7d82268b94 |
| ENCFF785IPD.tabix.bed.gz.tbi | 154.95 KB | 59f42ee872dece853e6276ea2ffc2f41 |
| ENCFF785IPD.tabix.bed.gz.tbi.dvc | 110.0 B | db37fada663ac681130f0c548b49e03e |
| genomic_resource.yaml | 2.03 KB | efff230e59334a387c4fb27c12951553 |
| genomic_resource_original.yaml | 1.9 KB | cfd71af67b7ddac0f6646eafbea21b91 |
| statistics/ |