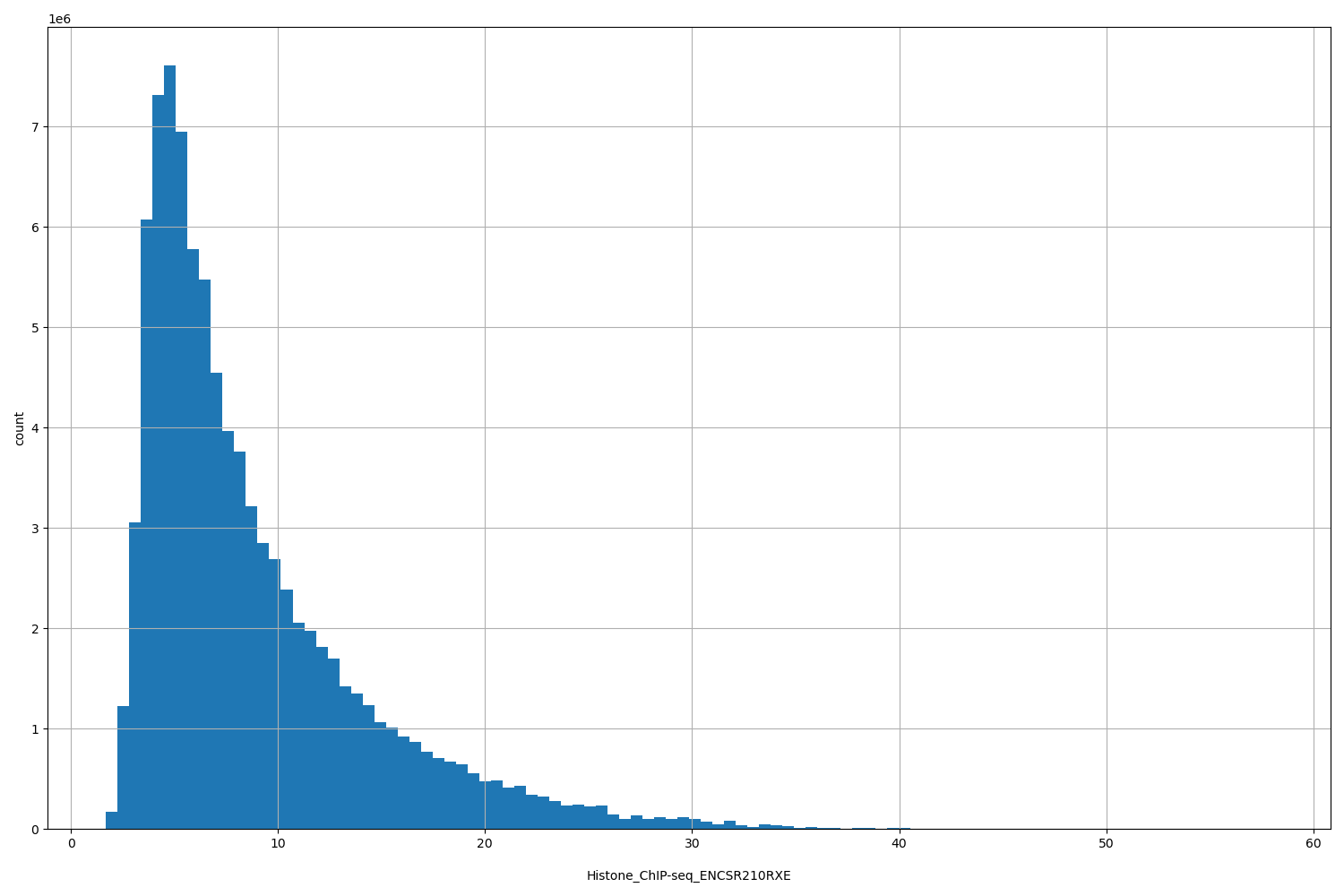
Histone_ChIP-seq_ENCSR210RXE
| Id: | Histone_ChIP-seq/ENCSR210RXE |
| Type: | position_score |
| Version: | 0 |
| Summary: |
HistoneChIP-seq ENCSR210RXE [biosamplesummary="Homo sapiens IMR-90" and target="H2BK120ac"] |
| Description: |
status: released biological_replicates: Rep 1, Rep 2 summary: output_type: replicated peaks audit_internal_action: Archived analysis {ENCAN283ALK|/analyses/ENCAN283ALK/} has in progress subobject quality standard {encode3-histone-chip|/quality-standards/encode3-histone-chip/} audit_warning: Processed alignments file {ENCFF997TDI|/files/ENCFF997TDI/} processed by ChIP-seq ENCODE3 hg19 pipeline has 11333713 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting H2BK120ac-human and investigated as a transcription factor is 10 million usable fragments. The recommended value is > 20 million, but > 10 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/} ) |
| Labels: |
|
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| Histone_ChIP-seq_ENCSR210RXE | float |
Histone_ChIP-seq_ENCSR210RXE |
Histone_ChIP-seq ENCSR210RXE [biosample_summary="Homo sapiens IMR-90" and target="H2BK120ac"]
|
 |
[1.69, 58] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF808ADB.bed.gz | 4.34 MB | 45dccfa558bc45b2760c27a9cc560b4a |
| ENCFF808ADB.bed.gz.dvc | 101.0 B | bf92518013d1ee12971683c51678d01a |
| ENCFF808ADB.tabix.bed.gz | 1.88 MB | bf944c218373d6e283299369ed978b50 |
| ENCFF808ADB.tabix.bed.gz.dvc | 107.0 B | 27edb1f1f50272d8ba813787c38b7157 |
| ENCFF808ADB.tabix.bed.gz.tbi | 300.05 KB | 831d9eb09061a1e2e78e63dd550ef221 |
| ENCFF808ADB.tabix.bed.gz.tbi.dvc | 110.0 B | ba8f5cf00f89092bd71fed67f288a04b |
| genomic_resource.yaml | 1.87 KB | 6b7458fe4e24f8008ad465acd5b683a6 |
| genomic_resource_original.yaml | 1.76 KB | 00a7d3d259779427336ba753ee86f545 |
| statistics/ |