Histone_ChIP-seq_ENCSR161HZJ
| Id: | Histone_ChIP-seq/ENCSR161HZJ |
| Type: | position_score |
| Version: | 0 |
| Summary: |
HistoneChIP-seq ENCSR161HZJ [biosamplesummary="Homo sapiens gastrocnemius medialis tissue male adult (54 years)" and target="H3K4me1"] |
| Description: |
status: released biological_replicates: Rep 1, Rep 2 summary: male adult (54 years) output_type: pseudoreplicated peaks audit_internal_action: Released analysis {ENCAN774UTC|/analyses/ENCAN774UTC/} has in progress subobject document {2f3eec46-15c8-489a-b052-8a94ab6410b2|/documents/2f3eec46-15c8-489a-b052-8a94ab6410b2/} audit_internal_action: Released analysis {ENCAN774UTC|/analyses/ENCAN774UTC/} has in progress subobject quality standard {encode4-histone-chip|/quality-standards/encode4-histone-chip/} audit_not_compliant: Processed alignments file {ENCFF570RLH|/files/ENCFF570RLH/} processed by ChIP-seq ENCODE4 v1.5.1 GRCh38 pipeline has 18535269 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting H3K4me1-human and investigated as a broad histone mark is 20 million usable fragments. The recommended value is > 45 million, but > 35 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/}) audit_not_compliant: Processed alignments file {ENCFF938FTW|/files/ENCFF938FTW/} processed by ChIP-seq ENCODE4 v1.5.1 GRCh38 pipeline has 33066155 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting H3K4me1-human and investigated as a broad histone mark is 20 million usable fragments. The recommended value is > 45 million, but > 35 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/}) |
| Labels: |
|
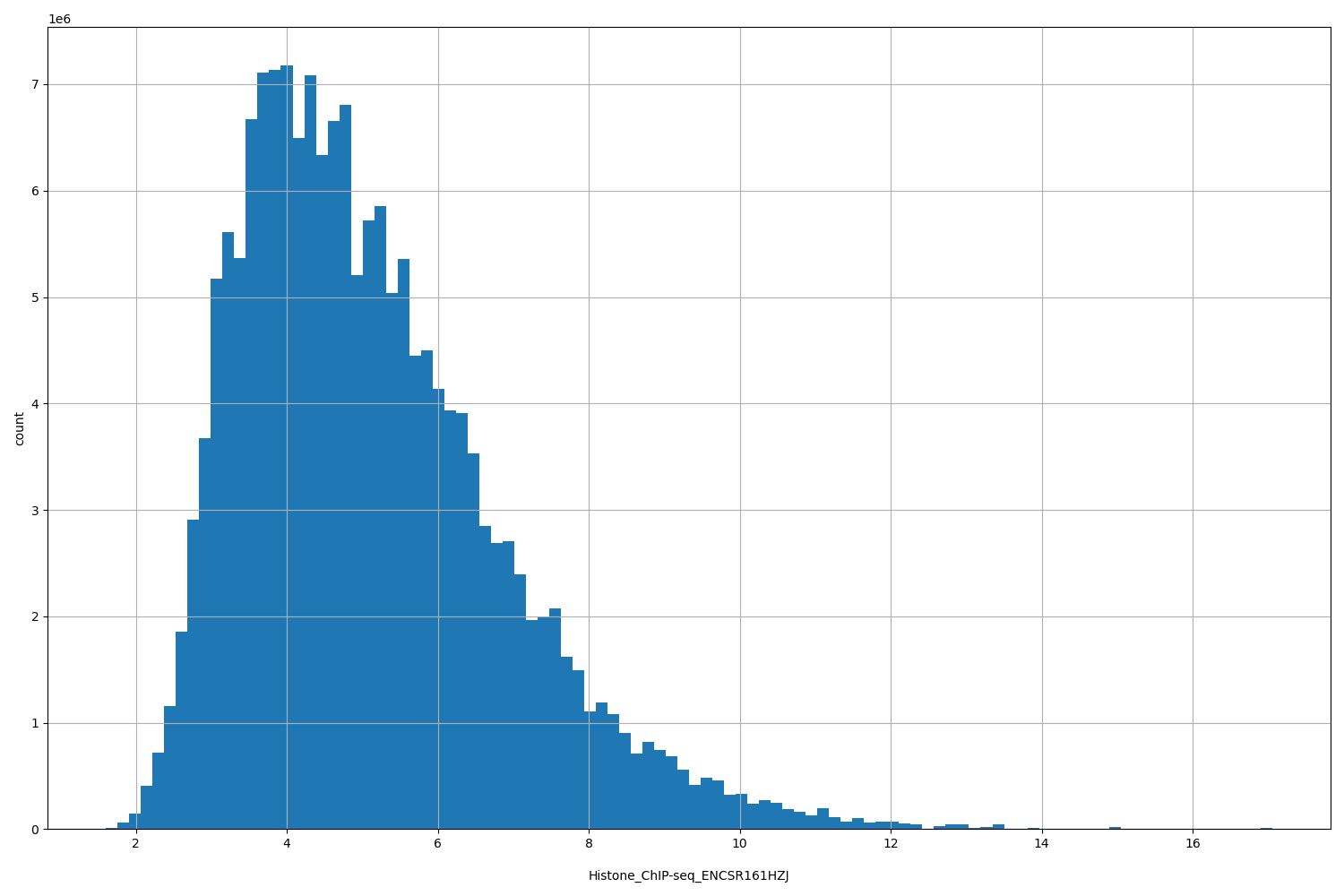
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| Histone_ChIP-seq_ENCSR161HZJ | float |
Histone_ChIP-seq_ENCSR161HZJ |
Histone_ChIP-seq ENCSR161HZJ [biosample_summary="Homo sapiens gastrocnemius medialis tissue male adult (54 years)" and target="H3K4me1"]
|
 |
[1.6, 17.1] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF574XMB.bed.gz | 2.84 MB | 41a76380c32a71fa5a659ec1fb681eaa |
| ENCFF574XMB.bed.gz.dvc | 101.0 B | 408c1fdf55de3b9f1604b2c7218acaae |
| ENCFF574XMB.tabix.bed.gz | 1.27 MB | 4993001093e36a10bda8f6ccffbd1062 |
| ENCFF574XMB.tabix.bed.gz.dvc | 107.0 B | 5a6025a06d89f4d215f611bccaf6020d |
| ENCFF574XMB.tabix.bed.gz.tbi | 287.1 KB | 6a224d45f93ec7afc708bce5fc20d84c |
| ENCFF574XMB.tabix.bed.gz.tbi.dvc | 110.0 B | 71c554f4ab14a5301d5848510ce1b080 |
| genomic_resource.yaml | 2.73 KB | cc32e53e02936ffaec9bb789b222c6ff |
| genomic_resource_original.yaml | 2.58 KB | aab1c485d57f36a1db3c1df4bf3fb83b |
| statistics/ |