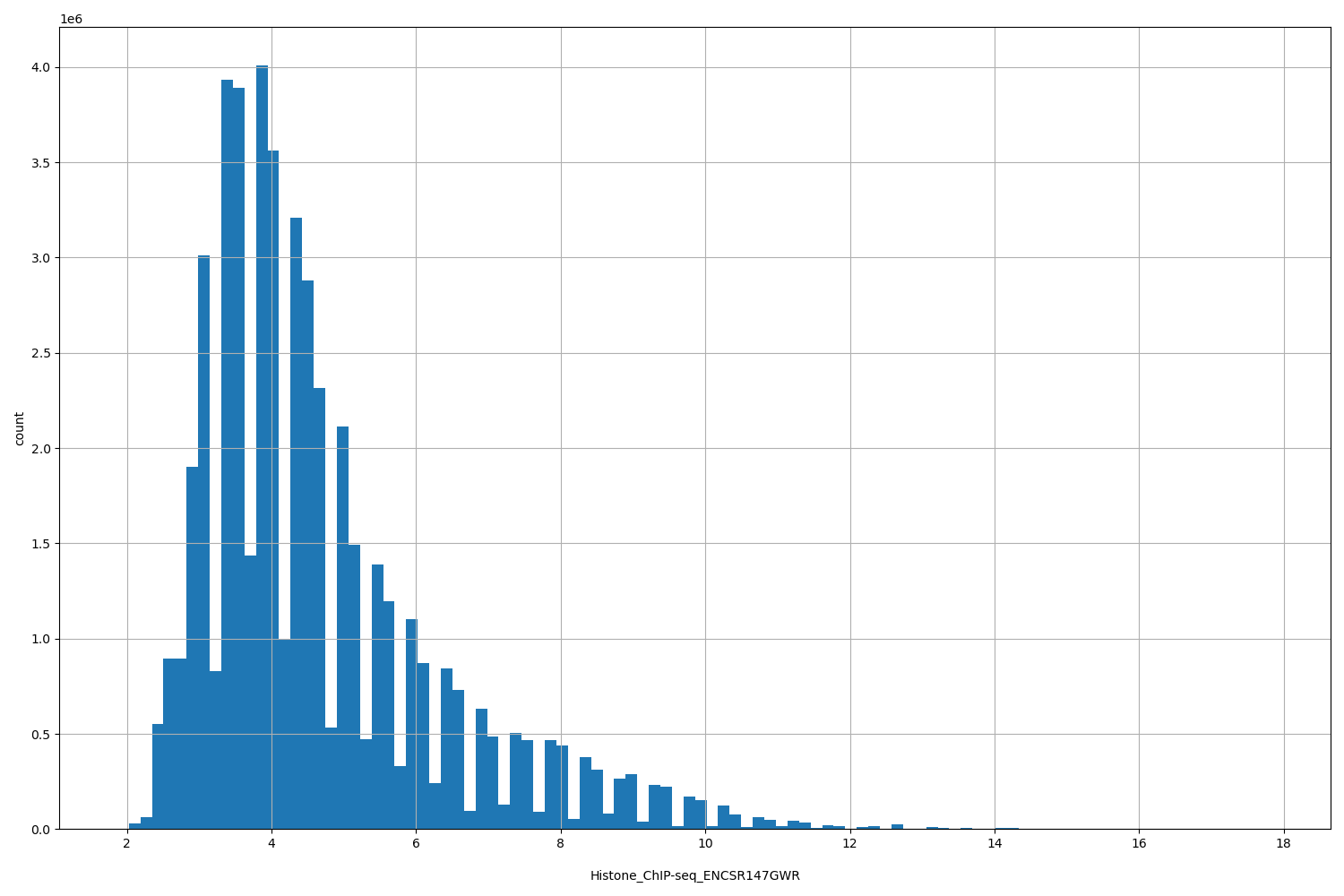
Histone_ChIP-seq_ENCSR147GWR
| Id: | Histone_ChIP-seq/ENCSR147GWR |
| Type: | position_score |
| Version: | 0 |
| Summary: |
HistoneChIP-seq ENCSR147GWR [biosamplesummary="Homo sapiens foreskin melanocyte male newborn" and target="H3K27me3"] |
| Description: |
status: released biological_replicates: Rep 1 summary: male newborn output_type: pseudoreplicated peaks audit_internal_action: Released analysis {ENCAN052PPP|/analyses/ENCAN052PPP/} has in progress subobject quality standard {encode4-histone-chip|/quality-standards/encode4-histone-chip/} audit_warning: Processed redacted alignments file {ENCFF640JPZ|/files/ENCFF640JPZ/} processed by ChIP-seq ENCODE4 v1.7.0 GRCh38 pipeline has 38653854 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting H3K27me3-human and investigated as a broad histone mark is 20 million usable fragments. The recommended value is > 45 million, but > 35 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/}) |
| Labels: |
|
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| Histone_ChIP-seq_ENCSR147GWR | float |
Histone_ChIP-seq_ENCSR147GWR |
Histone_ChIP-seq ENCSR147GWR [biosample_summary="Homo sapiens foreskin melanocyte male newborn" and target="H3K27me3"]
|
 |
[1.87, 17.8] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF250MQB.bed.gz | 1.91 MB | a13f697b12a045cccdfb6b97a3b8b323 |
| ENCFF250MQB.bed.gz.dvc | 101.0 B | 494ac34587328c23ca85b9a33980cfa6 |
| ENCFF250MQB.tabix.bed.gz | 1.03 MB | 1da07d621791281f83acdacc5773cf15 |
| ENCFF250MQB.tabix.bed.gz.dvc | 107.0 B | 1dce2337ede9c3dab04c56350f62b56d |
| ENCFF250MQB.tabix.bed.gz.tbi | 254.46 KB | 88f9f9591297d2875394646347f186fe |
| ENCFF250MQB.tabix.bed.gz.tbi.dvc | 110.0 B | bd079abd49742ec34f0f4abe59dd669a |
| genomic_resource.yaml | 1.96 KB | e000670bb91e16c42b04c29f3b33c589 |
| genomic_resource_original.yaml | 1.83 KB | 103bda58165a084dd14a56cf1bfc22ae |
| statistics/ |