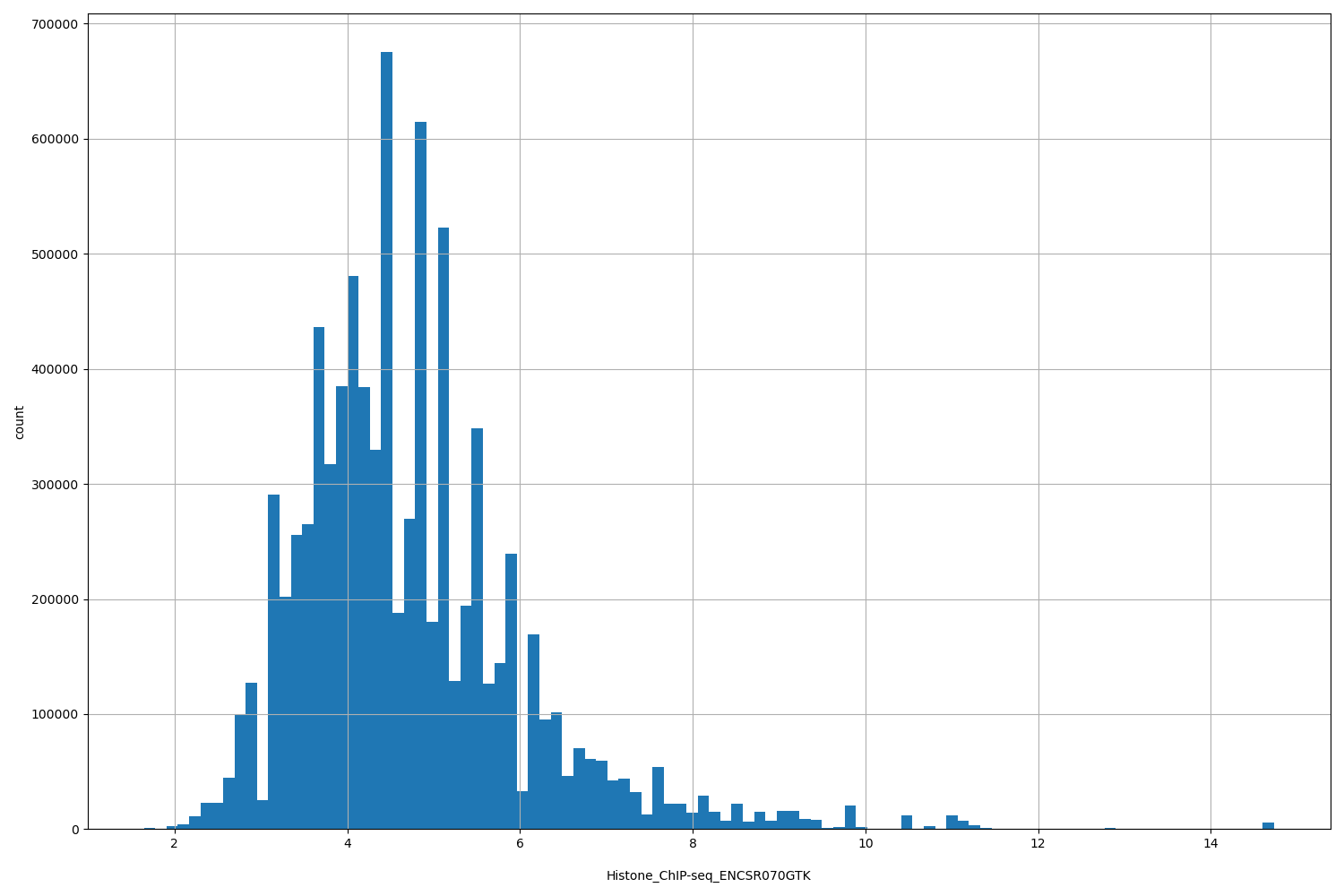
Histone_ChIP-seq_ENCSR070GTK
| Id: | Histone_ChIP-seq/ENCSR070GTK |
| Type: | position_score |
| Version: | 0 |
| Summary: |
HistoneChIP-seq ENCSR070GTK [biosamplesummary="Homo sapiens cingulate gyrus tissue male adult (81 years)" and target="H3K9me3"] |
| Description: |
status: released biological_replicates: Rep 1 summary: male adult (81 years) output_type: pseudoreplicated peaks audit_internal_action: Archived analysis {ENCAN250FCJ|/analyses/ENCAN250FCJ/} has in progress subobject quality standard {encode3-histone-chip|/quality-standards/encode3-histone-chip/} audit_not_compliant: Processed redacted unfiltered alignments file {ENCFF659URT|/files/ENCFF659URT/} processed by ChIP-seq ENCODE3 hg19 pipeline has 29200046 mapped reads. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting H3K9me3-human and investigated as a broad histone mark is 35 million mapped reads. The recommended value is > 45 million, but > 35 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/}) |
| Labels: |
|
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| Histone_ChIP-seq_ENCSR070GTK | float |
Histone_ChIP-seq_ENCSR070GTK |
Histone_ChIP-seq ENCSR070GTK [biosample_summary="Homo sapiens cingulate gyrus tissue male adult (81 years)" and target="H3K9me3"]
|
 |
[1.65, 14.7] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF057AAX.bed.gz | 249.01 KB | 987374912d93dfba531d4fdc679adc2b |
| ENCFF057AAX.bed.gz.dvc | 100.0 B | 7ba920cb85659cffe451f3c22331e2aa |
| ENCFF057AAX.tabix.bed.gz | 130.49 KB | a114fef0fb8ec8322d28e48777f6ced1 |
| ENCFF057AAX.tabix.bed.gz.dvc | 106.0 B | 2f40f093bd2397563cc44f551d5985fc |
| ENCFF057AAX.tabix.bed.gz.tbi | 92.54 KB | 7653f87d73ab208aaff5d07283db2a7c |
| ENCFF057AAX.tabix.bed.gz.tbi.dvc | 109.0 B | d0c7162c1b84c9add2deb57f73ee92e0 |
| genomic_resource.yaml | 2.0 KB | cae1f9ef77c756b30f3c03cec6f21084 |
| genomic_resource_original.yaml | 1.86 KB | 0a6acd856de5adf1c10a500b5e3f277f |
| statistics/ |