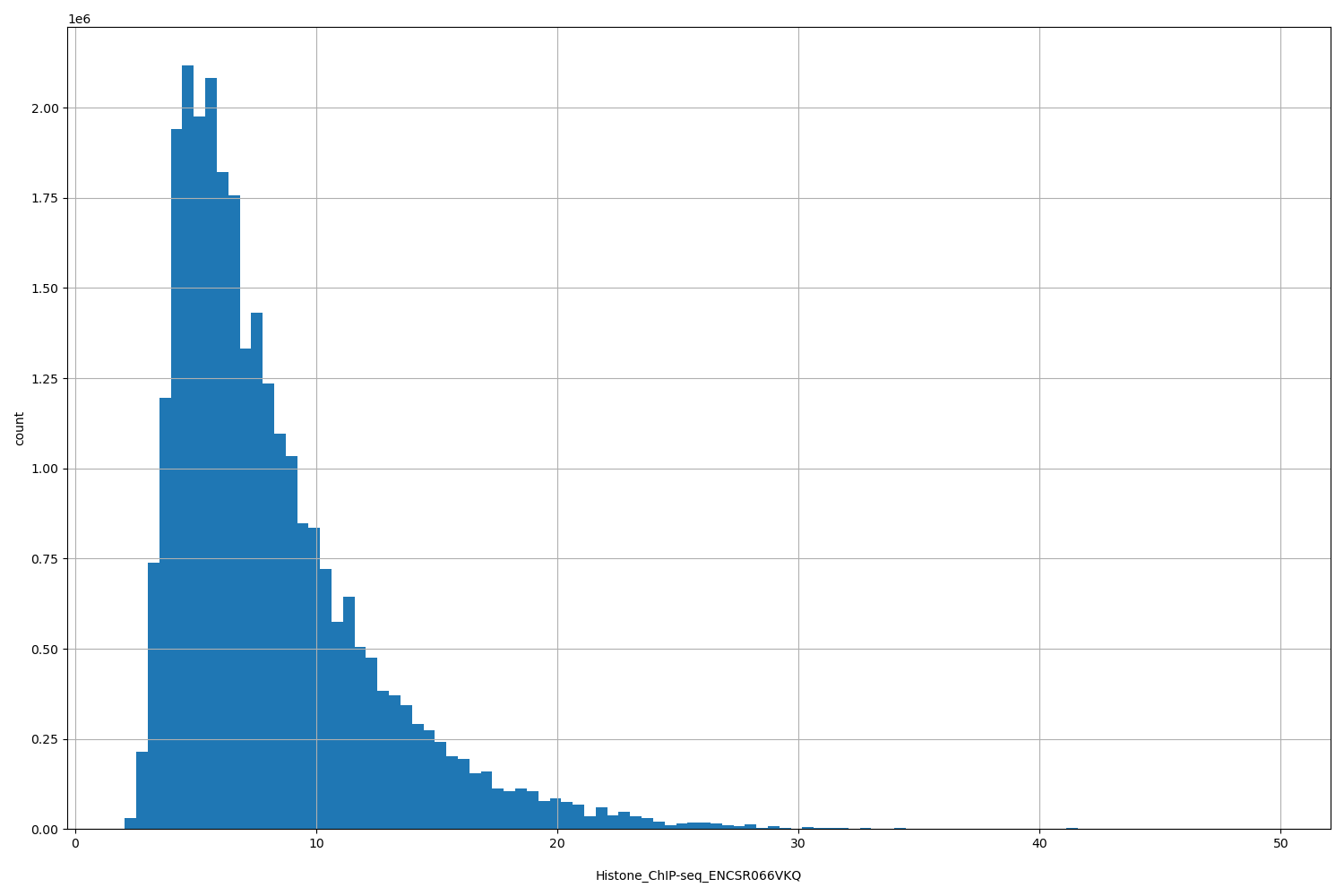
Histone_ChIP-seq_ENCSR066VKQ
| Id: | Histone_ChIP-seq/ENCSR066VKQ |
| Type: | position_score |
| Version: | 0 |
| Summary: |
HistoneChIP-seq ENCSR066VKQ [biosamplesummary="Homo sapiens IMR-90" and target="H3K56ac"] |
| Description: |
status: released biological_replicates: Rep 1, Rep 2 summary: output_type: pseudoreplicated peaks audit_error: Processed alignments file {ENCFF492AKV|/files/ENCFF492AKV/} processed by ChIP-seq ENCODE4 v1.6.1 GRCh38 pipeline has 4607368 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting H3K56ac-human and investigated as a transcription factor is 10 million usable fragments. The recommended value is > 20 million, but > 10 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/} ) audit_internal_action: Released analysis {ENCAN633CKM|/analyses/ENCAN633CKM/} has in progress subobject quality standard {encode4-histone-chip|/quality-standards/encode4-histone-chip/} audit_internal_action: Released analysis {ENCAN633CKM|/analyses/ENCAN633CKM/} has in progress subobject document {0c310adf-d961-4e11-9d77-fe398572c336|/documents/0c310adf-d961-4e11-9d77-fe398572c336/} audit_not_compliant: Processed alignments file {ENCFF281BMK|/files/ENCFF281BMK/} processed by ChIP-seq ENCODE4 v1.6.1 GRCh38 pipeline has 9738608 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting H3K56ac-human and investigated as a transcription factor is 10 million usable fragments. The recommended value is > 20 million, but > 10 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/} ) |
| Labels: |
|
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| Histone_ChIP-seq_ENCSR066VKQ | float |
Histone_ChIP-seq_ENCSR066VKQ |
Histone_ChIP-seq ENCSR066VKQ [biosample_summary="Homo sapiens IMR-90" and target="H3K56ac"]
|
 |
[2.07, 49.7] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF421NDL.bed.gz | 1.43 MB | 178d048f405e026de08e00cf8d07660b |
| ENCFF421NDL.bed.gz.dvc | 101.0 B | 521864e82dce6b5d6d621807779a1ede |
| ENCFF421NDL.tabix.bed.gz | 654.88 KB | 6373a69be2da873a354310480523721f |
| ENCFF421NDL.tabix.bed.gz.dvc | 106.0 B | ecc805f03b2c9685d77eac8f0895fb69 |
| ENCFF421NDL.tabix.bed.gz.tbi | 210.93 KB | c946bb8a5be66f4d97ab0d3d7e7755a2 |
| ENCFF421NDL.tabix.bed.gz.tbi.dvc | 110.0 B | 2ba32e14ee1a26ea0c59ddfce4fb1a68 |
| genomic_resource.yaml | 2.6 KB | 9d0f675bed5702a6a78f618063d83c3c |
| genomic_resource_original.yaml | 2.5 KB | ea8618c438888e10fbc95a1bb5465b44 |
| statistics/ |