Histone_ChIP-seq_ENCSR034BEQ
| Id: | Histone_ChIP-seq/ENCSR034BEQ |
| Type: | position_score |
| Version: | 0 |
| Summary: |
HistoneChIP-seq ENCSR034BEQ [biosamplesummary="Homo sapiens hepatocyte originated from H9" and target="H3K36me3"] |
| Description: |
status: released biological_replicates: Rep 1, Rep 2 summary: female embryo (5 days) output_type: replicated peaks audit_internal_action: Released analysis {ENCAN392XNX|/analyses/ENCAN392XNX/} has in progress subobject quality standard {encode4-histone-chip|/quality-standards/encode4-histone-chip/} audit_internal_action: Released analysis {ENCAN392XNX|/analyses/ENCAN392XNX/} has in progress subobject document {6c9816ef-978e-435b-a9a6-343d18c80d43|/documents/6c9816ef-978e-435b-a9a6-343d18c80d43/} audit_not_compliant: Processed alignments file {ENCFF980ZRR|/files/ENCFF980ZRR/} processed by ChIP-seq ENCODE4 v1.5.1 GRCh38 pipeline has 17368711 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting H3K36me3-human and investigated as a broad histone mark is 20 million usable fragments. The recommended value is > 45 million, but > 35 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/}) audit_warning: Processed alignments file {ENCFF641URR|/files/ENCFF641URR/} processed by ChIP-seq ENCODE4 v1.5.1 GRCh38 pipeline has 36852263 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting H3K36me3-human and investigated as a broad histone mark is 20 million usable fragments. The recommended value is > 45 million, but > 35 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/}) |
| Labels: |
|
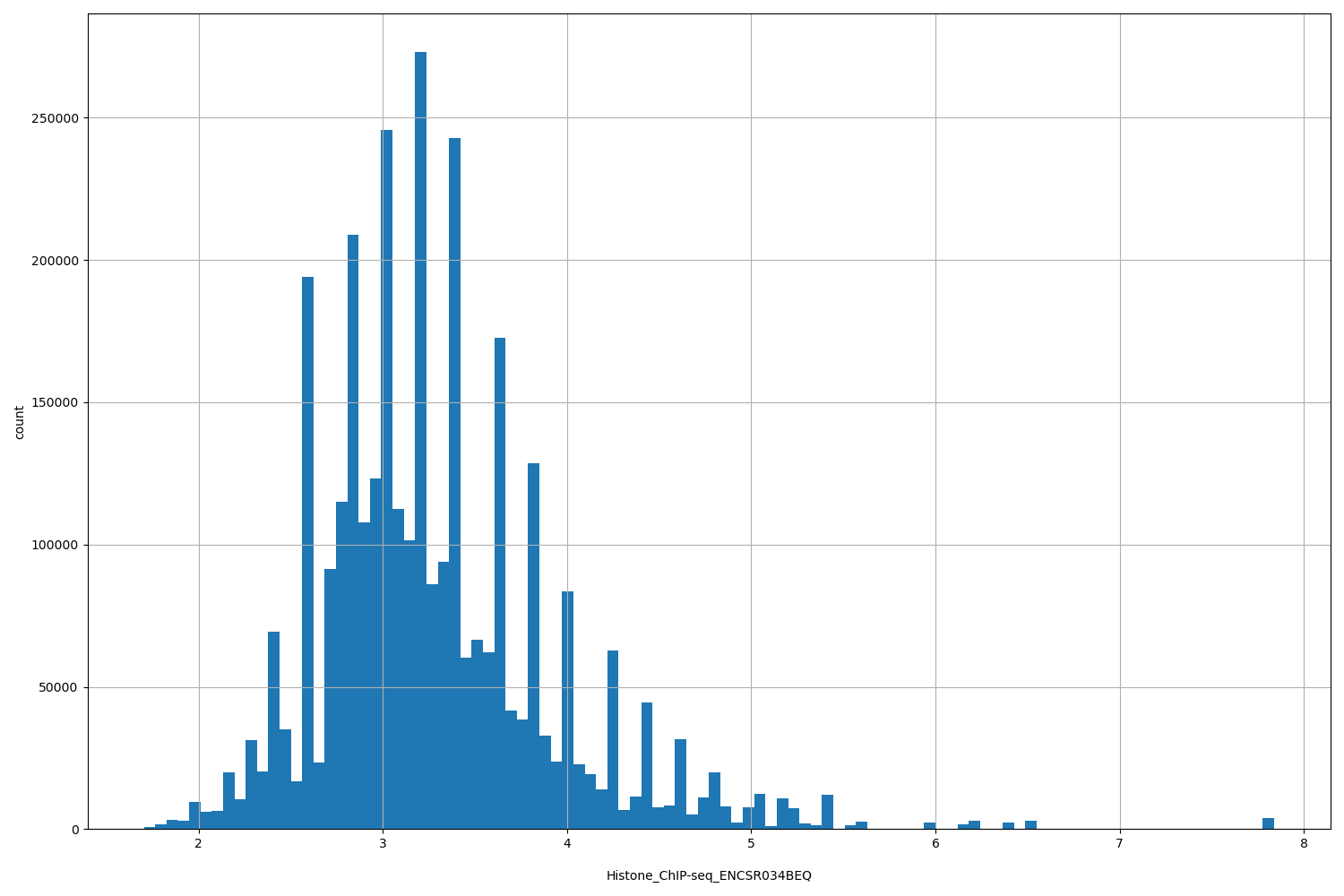
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| Histone_ChIP-seq_ENCSR034BEQ | float |
Histone_ChIP-seq_ENCSR034BEQ |
Histone_ChIP-seq ENCSR034BEQ [biosample_summary="Homo sapiens hepatocyte originated from H9" and target="H3K36me3"]
|
 |
[1.7, 7.84] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF980QVG.bed.gz | 123.99 KB | 62009c712041340475437d303ec3b1b1 |
| ENCFF980QVG.bed.gz.dvc | 100.0 B | fcaf45b2cfa5c14a5e64cef45eca91b2 |
| ENCFF980QVG.tabix.bed.gz | 64.16 KB | c5b7a402492e99be1d81d8f2c990d4c3 |
| ENCFF980QVG.tabix.bed.gz.dvc | 105.0 B | abe9c01d75d5379a76a0c7e535371f04 |
| ENCFF980QVG.tabix.bed.gz.tbi | 47.82 KB | e1572304b190c87889b2efe081280bc5 |
| ENCFF980QVG.tabix.bed.gz.tbi.dvc | 109.0 B | 5335391238fa7fe77b16ac942f8c2e15 |
| genomic_resource.yaml | 2.67 KB | bdb9f2c65ccf8739d70c0198ef8a6ccc |
| genomic_resource_original.yaml | 2.55 KB | a209247d1f67404f549fa7ebc2b06598 |
| statistics/ |