Histone_ChIP-seq_ENCSR031SHV
| Id: | Histone_ChIP-seq/ENCSR031SHV |
| Type: | position_score |
| Version: | 0 |
| Summary: |
HistoneChIP-seq ENCSR031SHV [biosamplesummary="Homo sapiens heart right ventricle tissue male adult (54 years)" and target="H3K4me3"] |
| Description: |
status: released biological_replicates: Rep 1 summary: male adult (54 years) output_type: pseudoreplicated peaks audit_internal_action: Released analysis {ENCAN764FLK|/analyses/ENCAN764FLK/} has in progress subobject quality standard {encode4-histone-chip|/quality-standards/encode4-histone-chip/} audit_warning: PBC1 (PCR Bottlenecking Coefficient 1, M1/M_distinct) is the ratio of the number of genomic locations where exactly one read maps uniquely (M1) to the number of genomic locations where some reads map (M_distinct). A PBC1 value in the range 0 - 0.5 is severe bottlenecking, 0.5 - 0.8 is moderate bottlenecking, 0.8 - 0.9 is mild bottlenecking, and > 0.9 is no bottlenecking. PBC1 value > 0.9 is recommended, but > 0.8 is acceptable. ENCODE processed alignments file {ENCFF802CWZ|/files/ENCFF802CWZ/} processed by ChIP-seq ENCODE4 v1.8.0 GRCh38 pipeline was generated from a library with PBC1 value of 0.84. audit_warning: PBC2 (PCR Bottlenecking Coefficient 2, M1/M2) is the ratio of the number of genomic locations where exactly one read maps uniquely (M1) to the number of genomic locations where two reads map uniquely (M2). A PBC2 value in the range 0 - 1 is severe bottlenecking, 1 - 3 is moderate bottlenecking, 3 - 10 is mild bottlenecking, > 10 is no bottlenecking. PBC2 value > 10 is recommended, but > 3 is acceptable. ENCODE processed alignments file {ENCFF802CWZ|/files/ENCFF802CWZ/} processed by ChIP-seq ENCODE4 v1.8.0 GRCh38 pipeline was generated from a library with PBC2 value of 7.12. |
| Labels: |
|
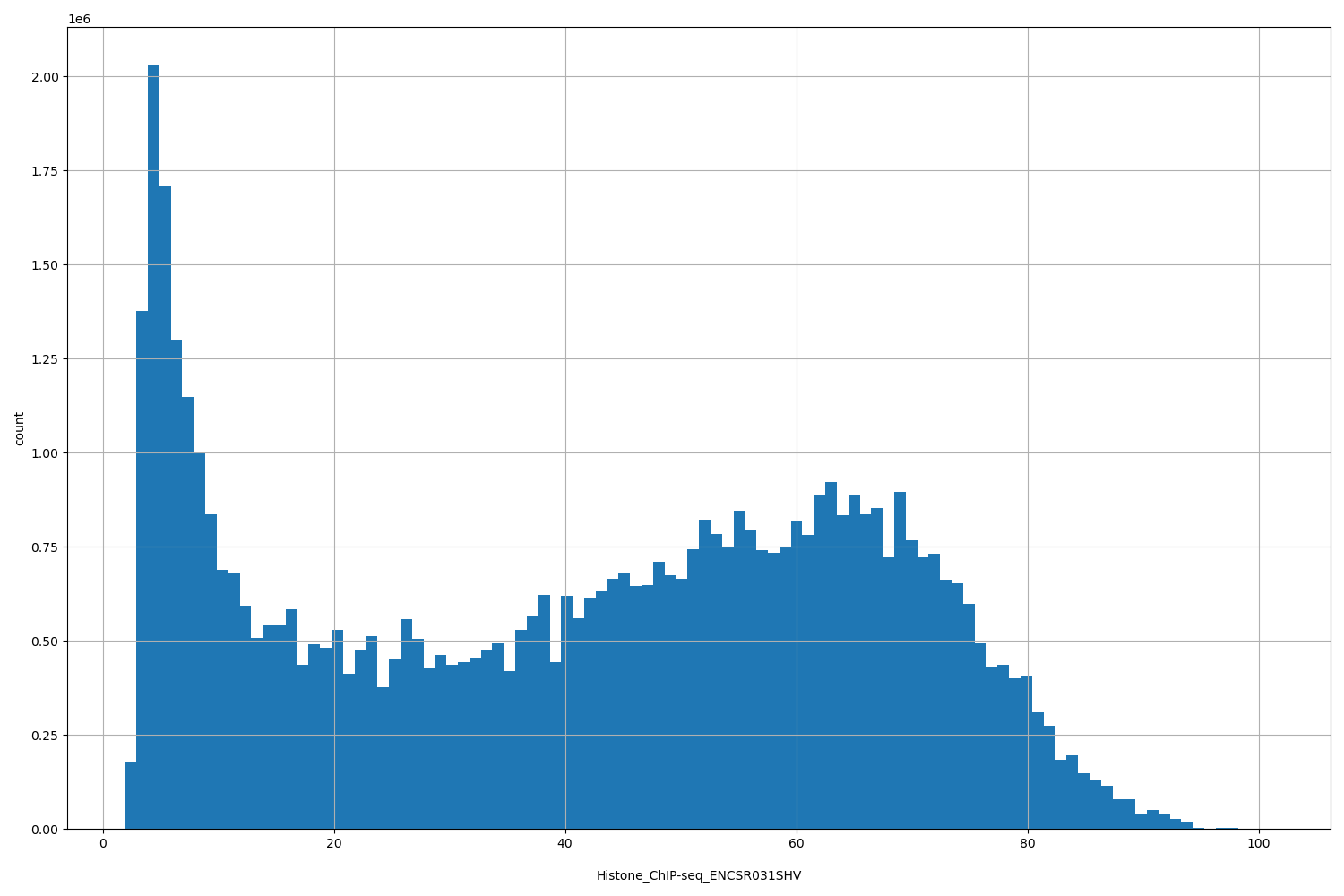
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| Histone_ChIP-seq_ENCSR031SHV | float |
Histone_ChIP-seq_ENCSR031SHV |
Histone_ChIP-seq ENCSR031SHV [biosample_summary="Homo sapiens heart right ventricle tissue male adult (54 years)" and target="H3K4me3"]
|
 |
[1.92, 101] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF577TDE.bed.gz | 840.56 KB | eff7798f4801e9811744404165c58434 |
| ENCFF577TDE.bed.gz.dvc | 100.0 B | 08fec8678d4793fb44d6da3e339dfc9f |
| ENCFF577TDE.tabix.bed.gz | 364.53 KB | adc4361e73bb0db7dadabc4479cdae38 |
| ENCFF577TDE.tabix.bed.gz.dvc | 106.0 B | e0498ee512c1bd5279df171fc8313702 |
| ENCFF577TDE.tabix.bed.gz.tbi | 192.3 KB | c54a3935a3879000711926bdd8b2e607 |
| ENCFF577TDE.tabix.bed.gz.tbi.dvc | 110.0 B | 0a07c251c50cc1910c29c84009dd3234 |
| genomic_resource.yaml | 2.79 KB | 2b33140f7ed1a75dde5ebfc361925663 |
| genomic_resource_original.yaml | 2.65 KB | 1dca42f38691e11e3275e67137a0b8c9 |
| statistics/ |