Histone_ChIP-seq_ENCSR021FSY
| Id: | Histone_ChIP-seq/ENCSR021FSY |
| Type: | position_score |
| Version: | 0 |
| Summary: |
HistoneChIP-seq ENCSR021FSY [biosamplesummary="Homo sapiens natural killer cell male adult (21 years)" and target="H3K9me3"] |
| Description: |
status: released biological_replicates: Rep 1 summary: male adult (21 years) output_type: pseudoreplicated peaks audit_internal_action: Archived analysis {ENCAN747WAG|/analyses/ENCAN747WAG/} has in progress subobject quality standard {encode3-histone-chip|/quality-standards/encode3-histone-chip/} audit_not_compliant: Processed unfiltered alignments file {ENCFF975BVW|/files/ENCFF975BVW/} processed by ChIP-seq ENCODE3 GRCh38 pipeline has 25743506 mapped reads. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting H3K9me3-human and investigated as a broad histone mark is 35 million mapped reads. The recommended value is > 45 million, but > 35 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/}) |
| Labels: |
|
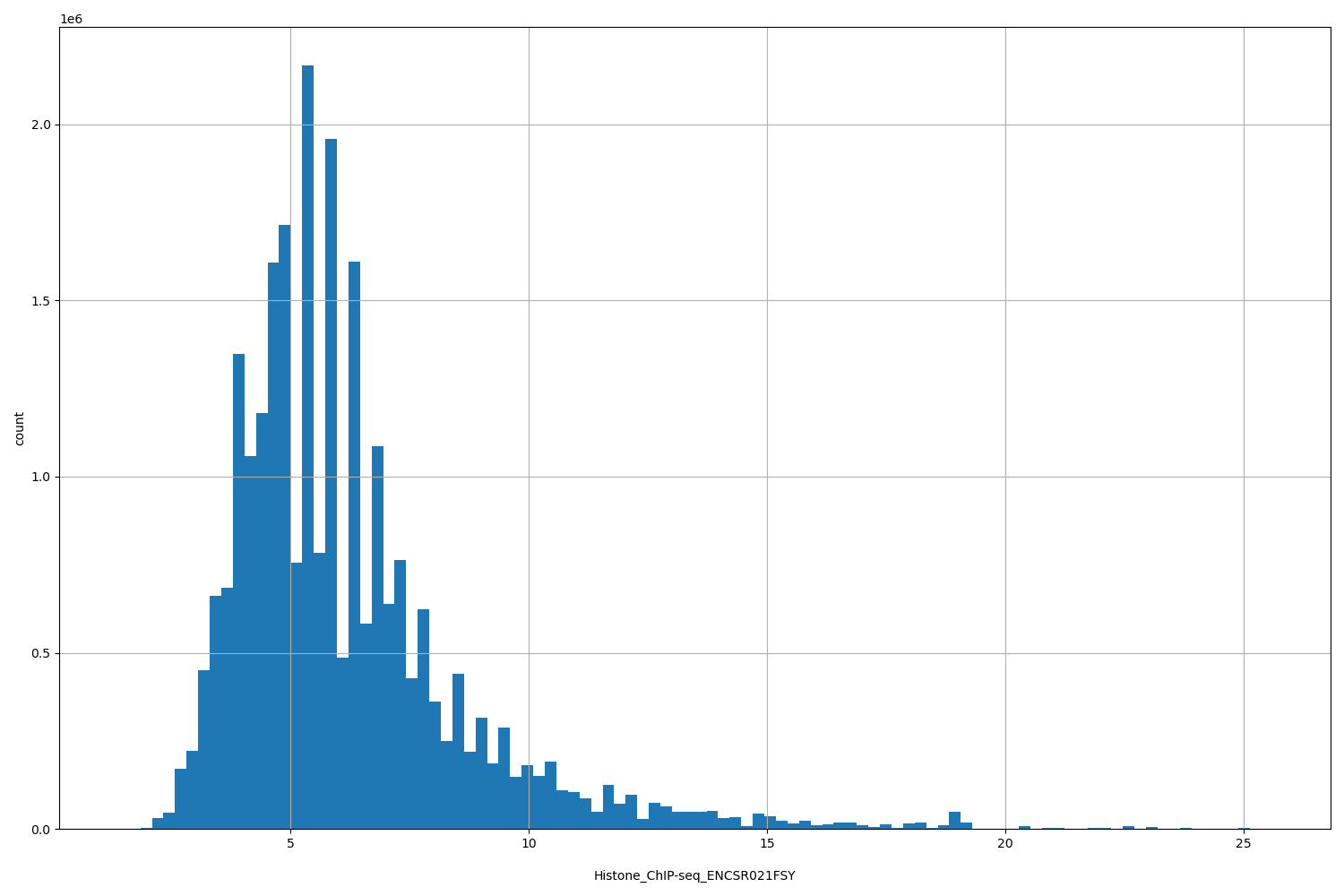
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| Histone_ChIP-seq_ENCSR021FSY | float |
Histone_ChIP-seq_ENCSR021FSY |
Histone_ChIP-seq ENCSR021FSY [biosample_summary="Homo sapiens natural killer cell male adult (21 years)" and target="H3K9me3"]
|
 |
[1.37, 25.6] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF622WDM.bed.gz | 936.66 KB | ea427f6a782f9e2e403bf715c36edc5e |
| ENCFF622WDM.bed.gz.dvc | 100.0 B | d5605ff17df1be674f1673d6bf4fe891 |
| ENCFF622WDM.tabix.bed.gz | 480.36 KB | 0d7d4bf8c1ec10486296b02336689114 |
| ENCFF622WDM.tabix.bed.gz.dvc | 106.0 B | 6276aec55622381fbd6caa945443415e |
| ENCFF622WDM.tabix.bed.gz.tbi | 236.59 KB | 9c8b9d86d9959a55d45d920e29e822c1 |
| ENCFF622WDM.tabix.bed.gz.tbi.dvc | 110.0 B | 64d285fedcbb1ee726cbe2fd950e2367 |
| genomic_resource.yaml | 1.99 KB | b9087e198063356528b389128cc97f1f |
| genomic_resource_original.yaml | 1.85 KB | 53e71a440187213a76c941c44cf4dca8 |
| statistics/ |