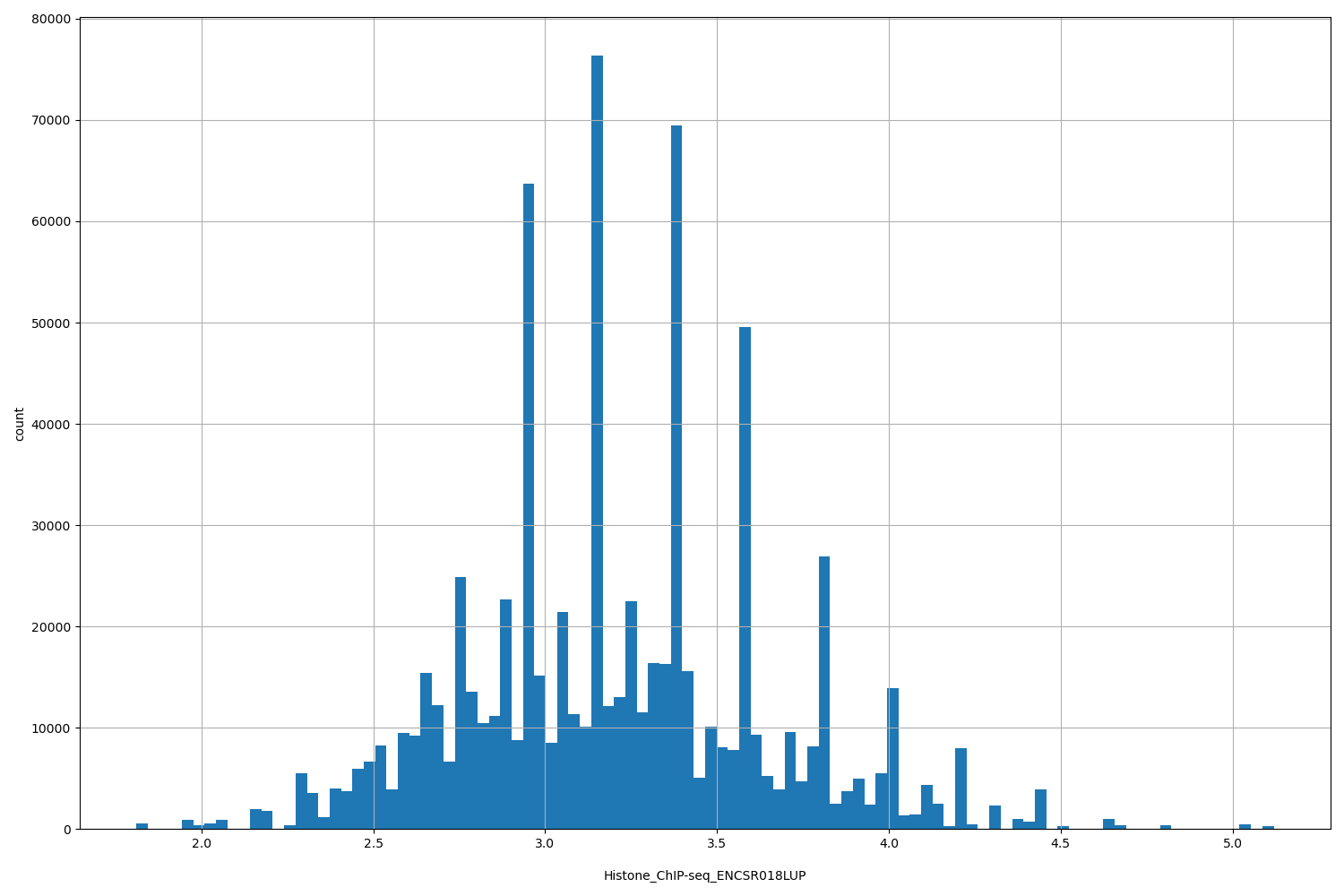
Histone_ChIP-seq_ENCSR018LUP
| Id: | Histone_ChIP-seq/ENCSR018LUP |
| Type: | position_score |
| Version: | 0 |
| Summary: |
HistoneChIP-seq ENCSR018LUP [biosamplesummary="Homo sapiens SU-DHL-6" and target="H3K9me2"] |
| Description: |
status: released biological_replicates: Rep 1, Rep 2 summary: output_type: replicated peaks audit_internal_action: Released analysis {ENCAN734NGS|/analyses/ENCAN734NGS/} has in progress subobject quality standard {encode4-histone-chip|/quality-standards/encode4-histone-chip/} audit_internal_action: Released analysis {ENCAN734NGS|/analyses/ENCAN734NGS/} has in progress subobject document {02111626-8295-4054-9188-5627c7ea91b7|/documents/02111626-8295-4054-9188-5627c7ea91b7/} audit_not_compliant: Processed alignments file {ENCFF617HNP|/files/ENCFF617HNP/} processed by ChIP-seq ENCODE4 v1.6.1 GRCh38 pipeline has 20076534 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting H3K9me2-human and investigated as a broad histone mark is 20 million usable fragments. The recommended value is > 45 million, but > 35 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/}) audit_not_compliant: Processed alignments file {ENCFF939ANP|/files/ENCFF939ANP/} processed by ChIP-seq ENCODE4 v1.6.1 GRCh38 pipeline has 28485122 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting H3K9me2-human and investigated as a broad histone mark is 20 million usable fragments. The recommended value is > 45 million, but > 35 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/}) |
| Labels: |
|
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| Histone_ChIP-seq_ENCSR018LUP | float |
Histone_ChIP-seq_ENCSR018LUP |
Histone_ChIP-seq ENCSR018LUP [biosample_summary="Homo sapiens SU-DHL-6" and target="H3K9me2"]
|
 |
[1.81, 5.12] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF929DUJ.bed.gz | 42.13 KB | 7148389f7388def2814df87b908175c7 |
| ENCFF929DUJ.bed.gz.dvc | 99.0 B | 5c194a9c261cf5a6399a487946fbf228 |
| ENCFF929DUJ.tabix.bed.gz | 21.99 KB | 28b29b79c74b743000be92823d21f02b |
| ENCFF929DUJ.tabix.bed.gz.dvc | 105.0 B | f4313ce981f786f0c80e4bdd6d737da7 |
| ENCFF929DUJ.tabix.bed.gz.tbi | 25.33 KB | c0ee31a9229584c20ea4f978a7752cce |
| ENCFF929DUJ.tabix.bed.gz.tbi.dvc | 109.0 B | 1c027010f225068cdbdf7a3570b5fafa |
| genomic_resource.yaml | 2.61 KB | 4a92662eebf9488ef4c39fb9eadbeea5 |
| genomic_resource_original.yaml | 2.5 KB | ad5137c9a7369099c7175c6f9acf28f1 |
| statistics/ |