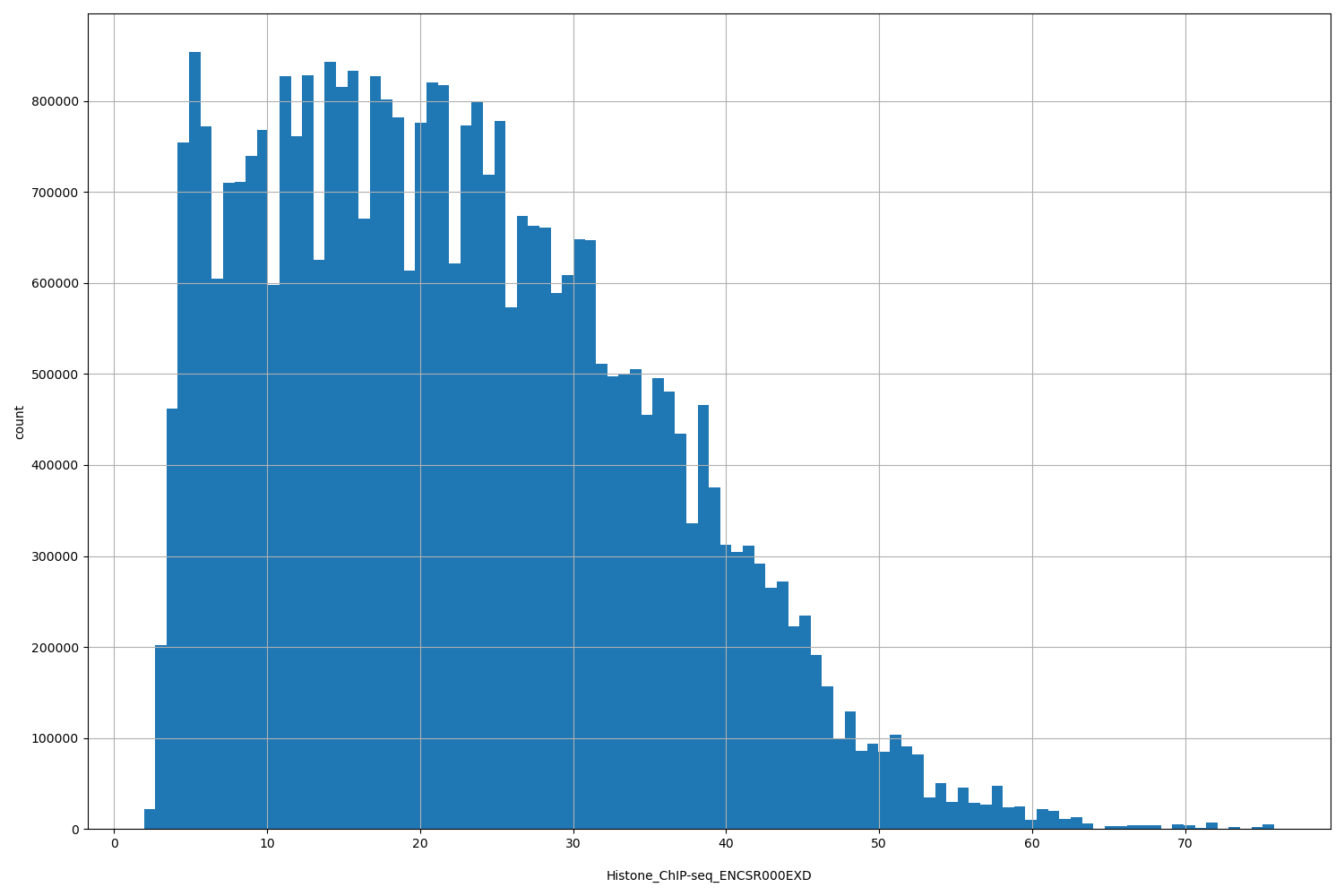
Histone_ChIP-seq_ENCSR000EXD
| Id: | Histone_ChIP-seq/ENCSR000EXD |
| Type: | position_score |
| Version: | 0 |
| Summary: |
HistoneChIP-seq ENCSR000EXD [biosamplesummary="Homo sapiens NT2/D1" and target="H3K4me3"] |
| Description: |
status: released biological_replicates: Rep 1, Rep 2 summary: output_type: pseudoreplicated peaks audit_internal_action: Released analysis {ENCAN494GVJ|/analyses/ENCAN494GVJ/} has in progress subobject quality standard {encode4-histone-chip|/quality-standards/encode4-histone-chip/} audit_internal_action: Released analysis {ENCAN494GVJ|/analyses/ENCAN494GVJ/} has in progress subobject document {aab2dde7-a679-4fb1-b12c-dfa499ce7b11|/documents/aab2dde7-a679-4fb1-b12c-dfa499ce7b11/} audit_warning: Processed alignments file {ENCFF595HXA|/files/ENCFF595HXA/} processed by ChIP-seq ENCODE4 v1.5.1 GRCh38 pipeline has 11633495 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting H3K4me3-human and investigated as a narrow histone mark is 10 million usable fragments. The recommended value is > 20 million, but > 10 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/}) audit_warning: Processed alignments file {ENCFF675KXV|/files/ENCFF675KXV/} processed by ChIP-seq ENCODE4 v1.5.1 GRCh38 pipeline has 12489051 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting H3K4me3-human and investigated as a narrow histone mark is 10 million usable fragments. The recommended value is > 20 million, but > 10 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/}) |
| Labels: |
|
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| Histone_ChIP-seq_ENCSR000EXD | float |
Histone_ChIP-seq_ENCSR000EXD |
Histone_ChIP-seq ENCSR000EXD [biosample_summary="Homo sapiens NT2/D1" and target="H3K4me3"]
|
 |
[1.95, 75.8] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF954FXD.bed.gz | 769.02 KB | e7c7678cfc765a96a6393513add730c6 |
| ENCFF954FXD.bed.gz.dvc | 100.0 B | e55ac428970474d088bab5bd65e312bb |
| ENCFF954FXD.tabix.bed.gz | 349.52 KB | 4b5af4012b288b4b38fd3e28cb3c46aa |
| ENCFF954FXD.tabix.bed.gz.dvc | 106.0 B | 025c1ad305531fc60999d8dd8a0dcdef |
| ENCFF954FXD.tabix.bed.gz.tbi | 157.68 KB | 9be831eda8058eccfbd0c53accb37bcb |
| ENCFF954FXD.tabix.bed.gz.tbi.dvc | 110.0 B | 712edcb6a309c10fc95d5613fcd0a193 |
| genomic_resource.yaml | 2.6 KB | 98ded614e2f584d6ffc801b56f50a4dd |
| genomic_resource_original.yaml | 2.49 KB | 25065a28a3364681f611da8613a4ab64 |
| statistics/ |