Histone_ChIP-seq_ENCSR000EXC
| Id: | Histone_ChIP-seq/ENCSR000EXC |
| Type: | position_score |
| Version: | 0 |
| Summary: |
HistoneChIP-seq ENCSR000EXC [biosamplesummary="Homo sapiens NT2/D1" and target="H3K9ac"] |
| Description: |
status: released biological_replicates: Rep 1, Rep 2 summary: output_type: replicated peaks audit_internal_action: Archived analysis {ENCAN423PFP|/analyses/ENCAN423PFP/} has in progress subobject quality standard {encode3-histone-chip|/quality-standards/encode3-histone-chip/} audit_warning: Processed alignments file {ENCFF144YZK|/files/ENCFF144YZK/} processed by ChIP-seq ENCODE3 hg19 pipeline has 11538761 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting H3K9ac-human and investigated as a narrow histone mark is 10 million usable fragments. The recommended value is > 20 million, but > 10 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/}) audit_warning: Processed alignments file {ENCFF493LSP|/files/ENCFF493LSP/} processed by ChIP-seq ENCODE3 hg19 pipeline has 12116599 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting H3K9ac-human and investigated as a narrow histone mark is 10 million usable fragments. The recommended value is > 20 million, but > 10 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/}) |
| Labels: |
|
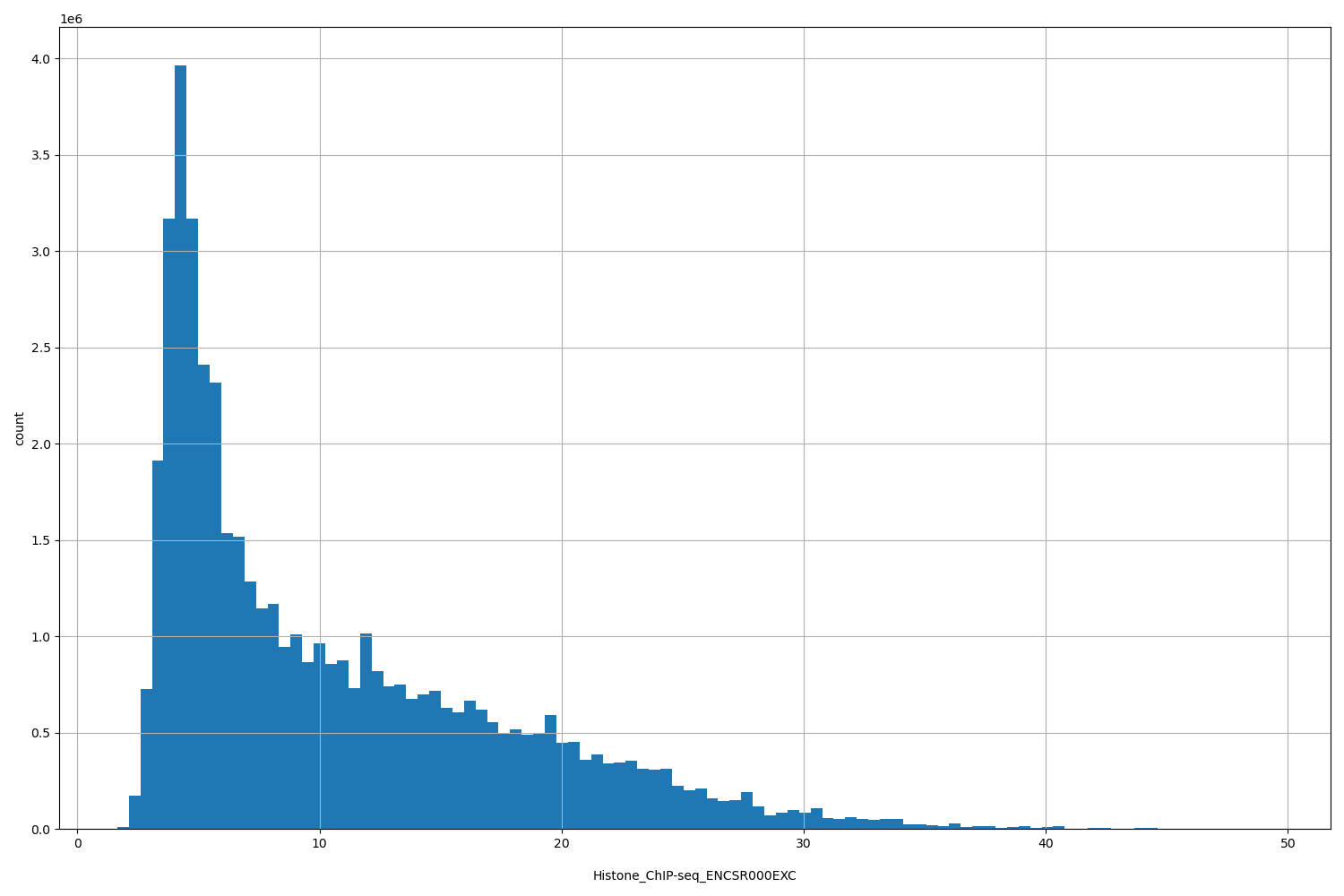
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| Histone_ChIP-seq_ENCSR000EXC | float |
Histone_ChIP-seq_ENCSR000EXC |
Histone_ChIP-seq ENCSR000EXC [biosample_summary="Homo sapiens NT2/D1" and target="H3K9ac"]
|
 |
[1.64, 49.4] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF304WSW.bed.gz | 1.66 MB | 7f15e7b7bf3bd96eed331f8a281aa463 |
| ENCFF304WSW.bed.gz.dvc | 101.0 B | 4cf5ac2e828a21d192421ac4f88f6338 |
| ENCFF304WSW.tabix.bed.gz | 793.12 KB | bdfc01638a979cfc2194c67feed3451e |
| ENCFF304WSW.tabix.bed.gz.dvc | 106.0 B | 3c222bba90e04ea1421fffcc574288c7 |
| ENCFF304WSW.tabix.bed.gz.tbi | 225.65 KB | 460687798feb95be48c1dda1c7d9dd29 |
| ENCFF304WSW.tabix.bed.gz.tbi.dvc | 110.0 B | cf2c7925b794d5371838d2e75f89e48a |
| genomic_resource.yaml | 2.35 KB | 8892bb8d539bf6ff7df4da6c299df4dc |
| genomic_resource_original.yaml | 2.24 KB | 683afa319605fb127e24a1e32ca0fc3b |
| statistics/ |