Histone_ChIP-seq_ENCSR000DXU
| Id: | Histone_ChIP-seq/ENCSR000DXU |
| Type: | position_score |
| Version: | 0 |
| Summary: |
HistoneChIP-seq ENCSR000DXU [biosamplesummary="Homo sapiens WERI-Rb-1" and target="H3K4me3"] |
| Description: |
status: released biological_replicates: Rep 1, Rep 2 summary: output_type: replicated peaks audit_internal_action: Archived analysis {ENCAN010DPE|/analyses/ENCAN010DPE/} has in progress subobject quality standard {encode3-histone-chip|/quality-standards/encode3-histone-chip/} audit_warning: Processed alignments file {ENCFF195NMV|/files/ENCFF195NMV/} processed by ChIP-seq ENCODE3 hg19 pipeline has 14614602 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting H3K4me3-human and investigated as a narrow histone mark is 10 million usable fragments. The recommended value is > 20 million, but > 10 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/}) audit_warning: Processed alignments file {ENCFF289SMU|/files/ENCFF289SMU/} processed by ChIP-seq ENCODE3 hg19 pipeline has 14897472 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting H3K4me3-human and investigated as a narrow histone mark is 10 million usable fragments. The recommended value is > 20 million, but > 10 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/}) audit_warning: PBC1 (PCR Bottlenecking Coefficient 1, M1/M_distinct) is the ratio of the number of genomic locations where exactly one read maps uniquely (M1) to the number of genomic locations where some reads map (M_distinct). A PBC1 value in the range 0 - 0.5 is severe bottlenecking, 0.5 - 0.8 is moderate bottlenecking, 0.8 - 0.9 is mild bottlenecking, and > 0.9 is no bottlenecking. PBC1 value > 0.9 is recommended, but > 0.8 is acceptable. ENCODE processed alignments file {ENCFF195NMV|/files/ENCFF195NMV/} processed by ChIP-seq ENCODE3 hg19 pipeline was generated from a library with PBC1 value of 0.89. audit_warning: PBC2 (PCR Bottlenecking Coefficient 2, M1/M2) is the ratio of the number of genomic locations where exactly one read maps uniquely (M1) to the number of genomic locations where two reads map uniquely (M2). A PBC2 value in the range 0 - 1 is severe bottlenecking, 1 - 3 is moderate bottlenecking, 3 - 10 is mild bottlenecking, > 10 is no bottlenecking. PBC2 value > 10 is recommended, but > 3 is acceptable. ENCODE processed alignments file {ENCFF195NMV|/files/ENCFF195NMV/} processed by ChIP-seq ENCODE3 hg19 pipeline was generated from a library with PBC2 value of 9.77. audit_warning: PBC1 (PCR Bottlenecking Coefficient 1, M1/M_distinct) is the ratio of the number of genomic locations where exactly one read maps uniquely (M1) to the number of genomic locations where some reads map (M_distinct). A PBC1 value in the range 0 - 0.5 is severe bottlenecking, 0.5 - 0.8 is moderate bottlenecking, 0.8 - 0.9 is mild bottlenecking, and > 0.9 is no bottlenecking. PBC1 value > 0.9 is recommended, but > 0.8 is acceptable. ENCODE processed alignments file {ENCFF289SMU|/files/ENCFF289SMU/} processed by ChIP-seq ENCODE3 hg19 pipeline was generated from a library with PBC1 value of 0.88. audit_warning: PBC2 (PCR Bottlenecking Coefficient 2, M1/M2) is the ratio of the number of genomic locations where exactly one read maps uniquely (M1) to the number of genomic locations where two reads map uniquely (M2). A PBC2 value in the range 0 - 1 is severe bottlenecking, 1 - 3 is moderate bottlenecking, 3 - 10 is mild bottlenecking, > 10 is no bottlenecking. PBC2 value > 10 is recommended, but > 3 is acceptable. ENCODE processed alignments file {ENCFF289SMU|/files/ENCFF289SMU/} processed by ChIP-seq ENCODE3 hg19 pipeline was generated from a library with PBC2 value of 8.88. |
| Labels: |
|
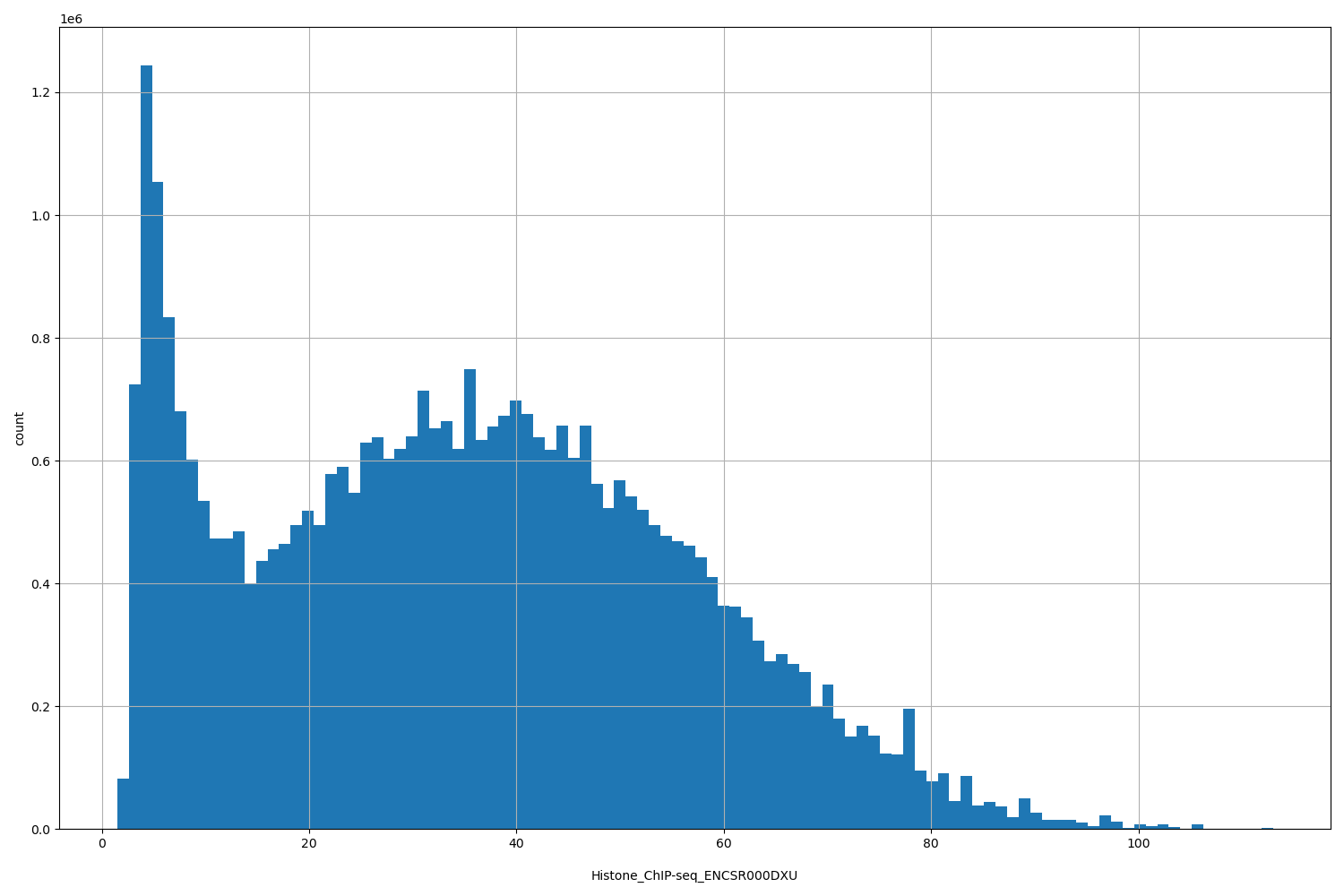
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| Histone_ChIP-seq_ENCSR000DXU | float |
Histone_ChIP-seq_ENCSR000DXU |
Histone_ChIP-seq ENCSR000DXU [biosample_summary="Homo sapiens WERI-Rb-1" and target="H3K4me3"]
|
 |
[1.49, 113] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF285MTN.bed.gz | 851.55 KB | bddda281fe9f4fd6de7495ff07a6a133 |
| ENCFF285MTN.bed.gz.dvc | 100.0 B | aa65e9742e4be713960268381e3f5064 |
| ENCFF285MTN.tabix.bed.gz | 362.18 KB | b3f6d3a949a00cec2e49e2dd7b2eeeaa |
| ENCFF285MTN.tabix.bed.gz.dvc | 106.0 B | 07ae8cfe06a25ded28cee226b5f76789 |
| ENCFF285MTN.tabix.bed.gz.tbi | 174.9 KB | 1743693871a3d6baabf3cb0c9864c43c |
| ENCFF285MTN.tabix.bed.gz.tbi.dvc | 110.0 B | d5a08d2f3ae58d444dc110eca806a29b |
| genomic_resource.yaml | 5.16 KB | 0f03f9bb76d9686e84e850d413a1a222 |
| genomic_resource_original.yaml | 5.05 KB | fb87d1480b9a368874560ec790b7977f |
| statistics/ |