Histone_ChIP-seq_ENCSR000DXK
| Id: | Histone_ChIP-seq/ENCSR000DXK |
| Type: | position_score |
| Version: | 0 |
| Summary: |
HistoneChIP-seq ENCSR000DXK [biosamplesummary="Homo sapiens bronchial epithelial cell" and target="H3K27me3"] |
| Description: |
status: released biological_replicates: Rep 1, Rep 2 summary: output_type: pseudoreplicated peaks audit_internal_action: Released analysis {ENCAN821XLN|/analyses/ENCAN821XLN/} has in progress subobject quality standard {encode4-histone-chip|/quality-standards/encode4-histone-chip/} audit_internal_action: Released analysis {ENCAN821XLN|/analyses/ENCAN821XLN/} has in progress subobject document {5cb7183b-9d3f-4f1c-9080-05034a5fce0b|/documents/5cb7183b-9d3f-4f1c-9080-05034a5fce0b/} audit_not_compliant: Processed alignments file {ENCFF024IIP|/files/ENCFF024IIP/} processed by ChIP-seq ENCODE4 v1.5.1 GRCh38 pipeline has 12215369 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting H3K27me3-human and investigated as a broad histone mark is 20 million usable fragments. The recommended value is > 45 million, but > 35 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/}) audit_not_compliant: Processed alignments file {ENCFF179MTQ|/files/ENCFF179MTQ/} processed by ChIP-seq ENCODE4 v1.5.1 GRCh38 pipeline has 13267302 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting H3K27me3-human and investigated as a broad histone mark is 20 million usable fragments. The recommended value is > 45 million, but > 35 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/}) |
| Labels: |
|
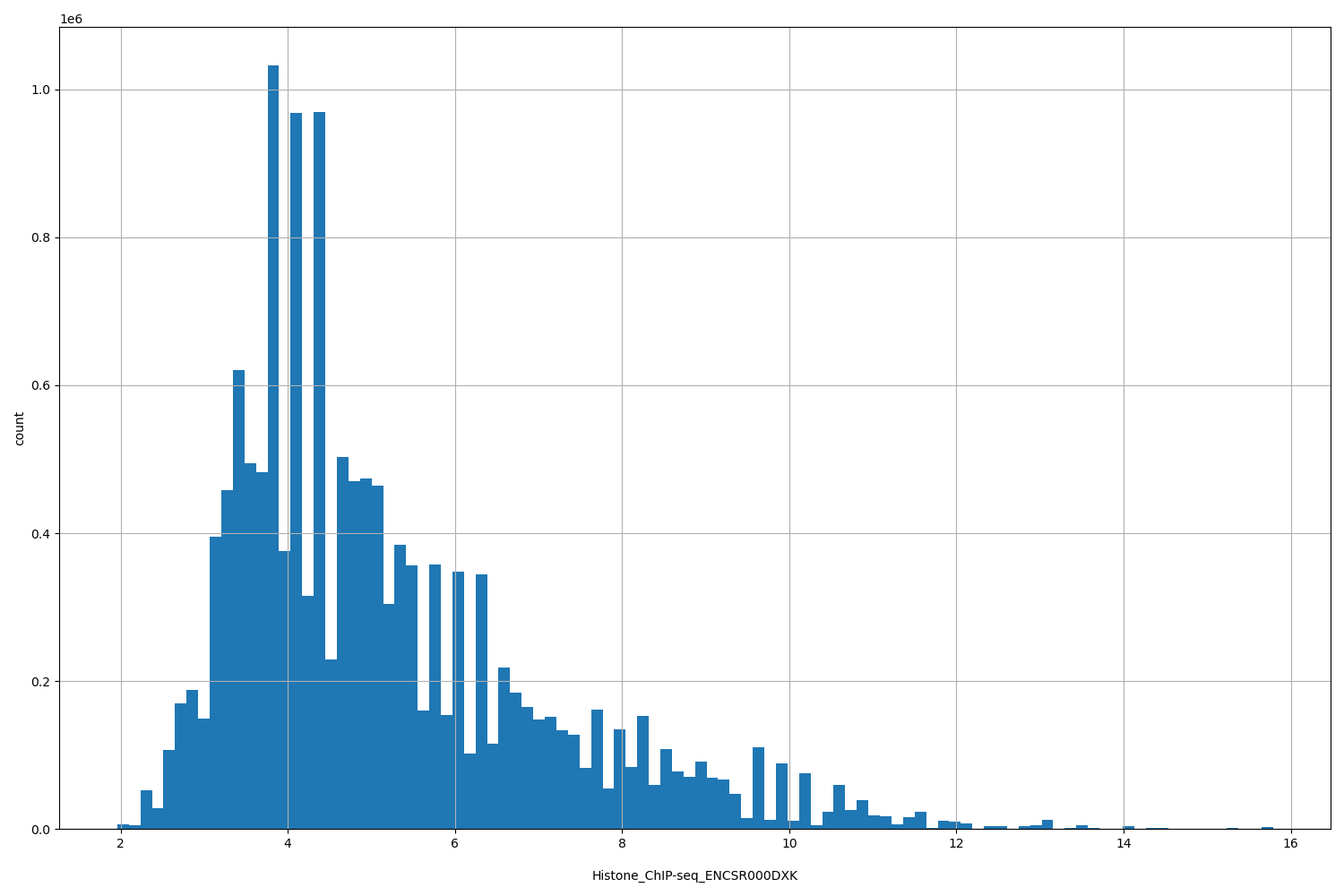
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| Histone_ChIP-seq_ENCSR000DXK | float |
Histone_ChIP-seq_ENCSR000DXK |
Histone_ChIP-seq ENCSR000DXK [biosample_summary="Homo sapiens bronchial epithelial cell" and target="H3K27me3"]
|
 |
[1.96, 15.8] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF495KNO.bed.gz | 787.14 KB | e2810fe7d18f1f6c7176260b00f25a8e |
| ENCFF495KNO.bed.gz.dvc | 100.0 B | a6c959d42e09af17e9a99ac5e34ac8c5 |
| ENCFF495KNO.tabix.bed.gz | 416.78 KB | f16f80958d11dbe79d170b4ca75e150c |
| ENCFF495KNO.tabix.bed.gz.dvc | 106.0 B | 67d0a983bf2c50a49cebbd96ac3bff73 |
| ENCFF495KNO.tabix.bed.gz.tbi | 141.86 KB | 771ac9bd1da369e51e79630191af3467 |
| ENCFF495KNO.tabix.bed.gz.tbi.dvc | 110.0 B | e62e02367141b38c5e2705713c37408b |
| genomic_resource.yaml | 2.97 KB | 0368781b576dd99c94ad996ee9e0b1a4 |
| genomic_resource_original.yaml | 2.85 KB | a1d0647c0525a30053558647403c5b52 |
| statistics/ |