Histone_ChIP-seq_ENCSR000DWM
| Id: | Histone_ChIP-seq/ENCSR000DWM |
| Type: | position_score |
| Version: | 0 |
| Summary: |
HistoneChIP-seq ENCSR000DWM [biosamplesummary="Homo sapiens CD14-positive monocyte female" and target="H3K27me3"] |
| Description: |
status: released biological_replicates: Rep 1 summary: female output_type: pseudoreplicated peaks audit_internal_action: Archived analysis {ENCAN564AMD|/analyses/ENCAN564AMD/} has in progress subobject quality standard {encode3-histone-chip|/quality-standards/encode3-histone-chip/} audit_not_compliant: Processed alignments file {ENCFF461EPJ|/files/ENCFF461EPJ/} processed by ChIP-seq ENCODE3 hg19 pipeline has 21354945 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting H3K27me3-human and investigated as a broad histone mark is 20 million usable fragments. The recommended value is > 45 million, but > 35 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/}) |
| Labels: |
|
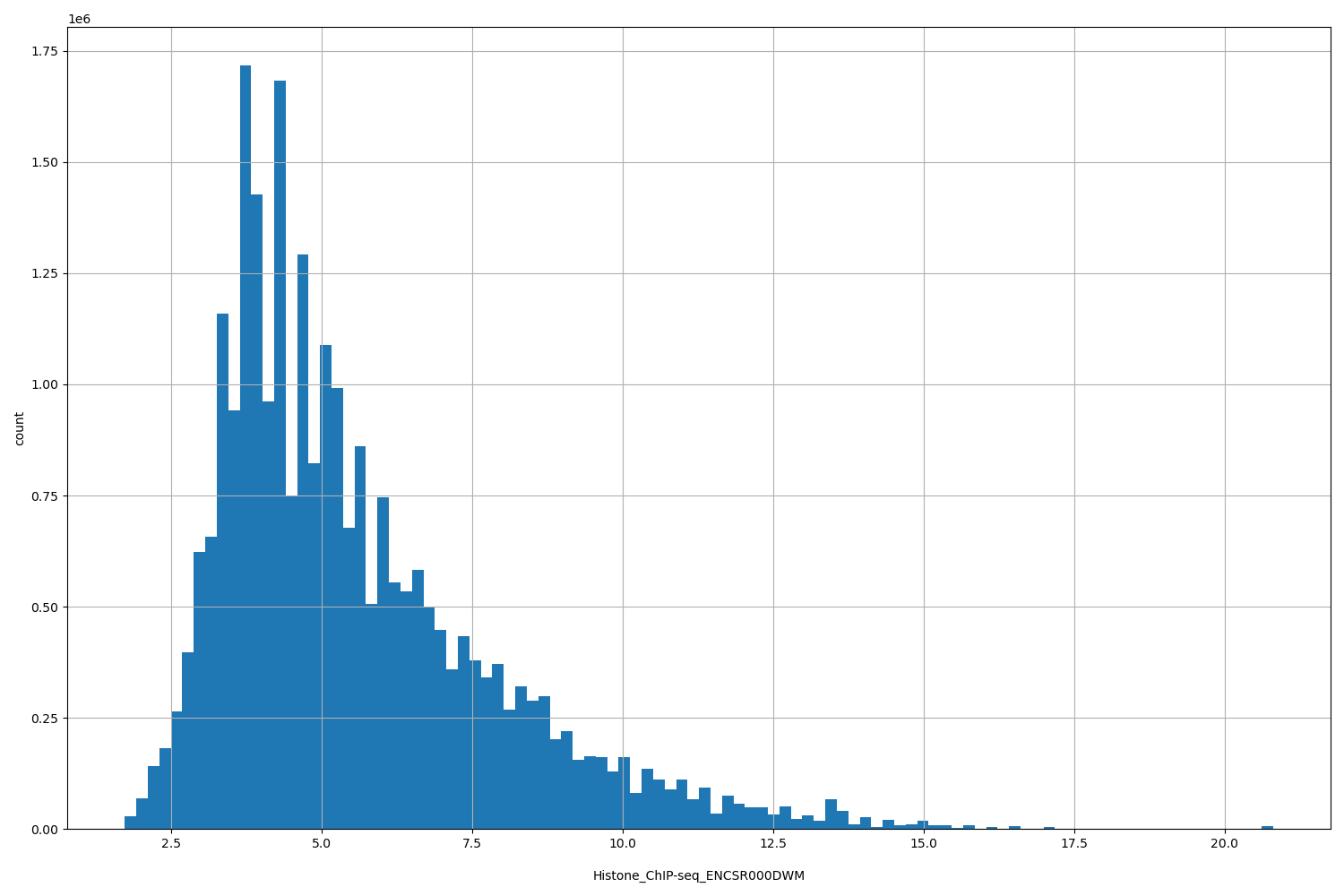
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| Histone_ChIP-seq_ENCSR000DWM | float |
Histone_ChIP-seq_ENCSR000DWM |
Histone_ChIP-seq ENCSR000DWM [biosample_summary="Homo sapiens CD14-positive monocyte female" and target="H3K27me3"]
|
 |
[1.73, 20.8] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF112ABQ.bed.gz | 1.01 MB | ef29cf55d02ce1a37473f99582ec7479 |
| ENCFF112ABQ.bed.gz.dvc | 101.0 B | cf4b943c5e99508509b680916e6fdcb3 |
| ENCFF112ABQ.tabix.bed.gz | 500.13 KB | 1986de0803de5708515086bd2f398e45 |
| ENCFF112ABQ.tabix.bed.gz.dvc | 106.0 B | fdcbd09c3670078b014be99864a79ade |
| ENCFF112ABQ.tabix.bed.gz.tbi | 123.65 KB | c50ac1fcc5a1ae9502185ff7852663df |
| ENCFF112ABQ.tabix.bed.gz.tbi.dvc | 110.0 B | 2e081f3996326079cc6f164ec689e0d7 |
| genomic_resource.yaml | 1.92 KB | 46a6d3da0043b5f314f57308902bbb58 |
| genomic_resource_original.yaml | 1.8 KB | 1fa90b60537c84268837da82e11e0898 |
| statistics/ |