Histone_ChIP-seq_ENCSR000DVO
| Id: | Histone_ChIP-seq/ENCSR000DVO |
| Type: | position_score |
| Version: | 0 |
| Summary: |
HistoneChIP-seq ENCSR000DVO [biosamplesummary="Homo sapiens endothelial cell of umbilical vein newborn" and target="H3K27me3"] |
| Description: |
status: released biological_replicates: Rep 1, Rep 2 summary: newborn output_type: replicated peaks audit_internal_action: Archived analysis {ENCAN710RBS|/analyses/ENCAN710RBS/} has in progress subobject quality standard {encode3-histone-chip|/quality-standards/encode3-histone-chip/} audit_not_compliant: Processed alignments file {ENCFF146ZZM|/files/ENCFF146ZZM/} processed by ChIP-seq ENCODE3 hg19 pipeline has 11696262 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting H3K27me3-human and investigated as a broad histone mark is 20 million usable fragments. The recommended value is > 45 million, but > 35 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/}) audit_not_compliant: Processed alignments file {ENCFF828NMX|/files/ENCFF828NMX/} processed by ChIP-seq ENCODE3 hg19 pipeline has 12538741 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting H3K27me3-human and investigated as a broad histone mark is 20 million usable fragments. The recommended value is > 45 million, but > 35 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/}) |
| Labels: |
|
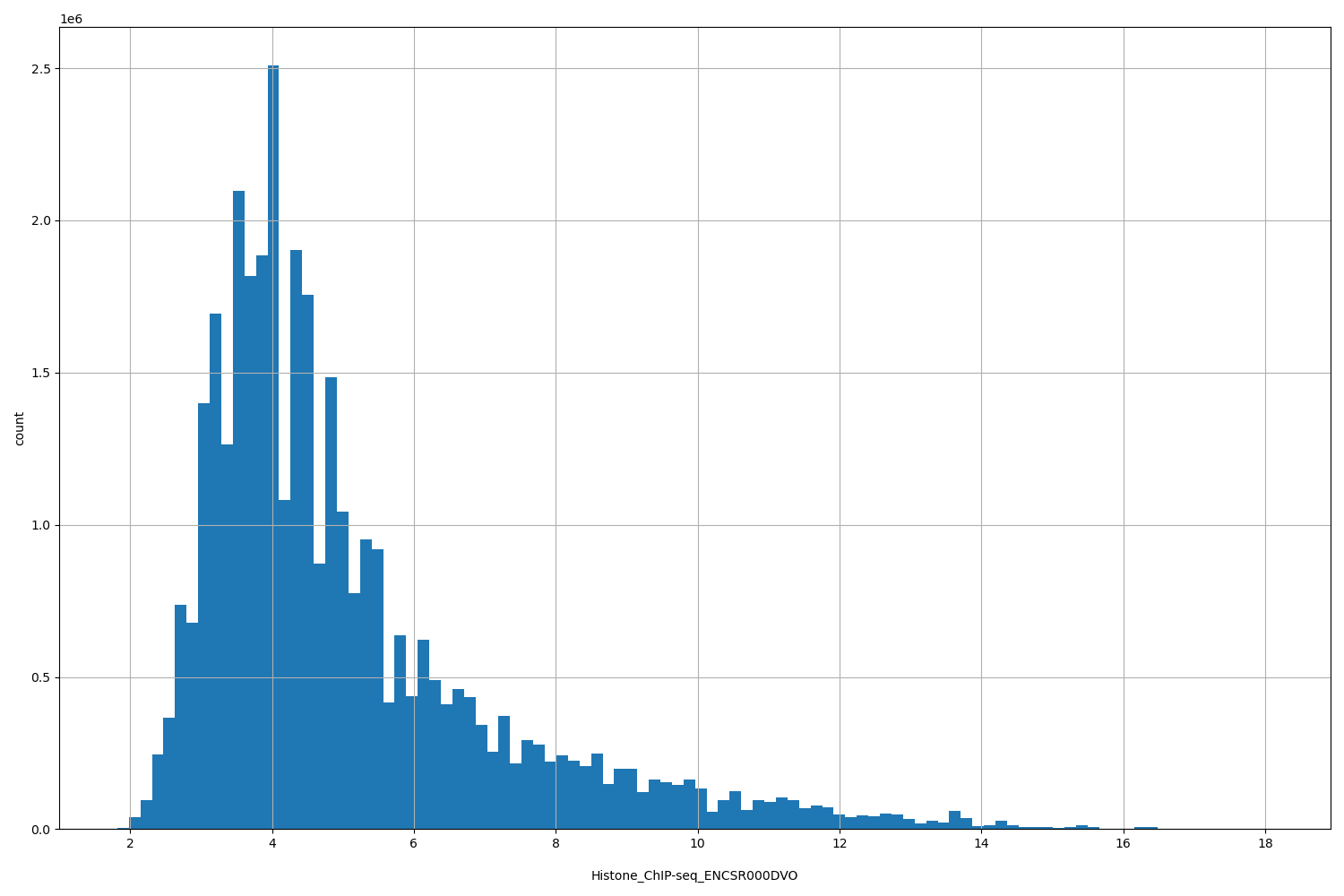
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| Histone_ChIP-seq_ENCSR000DVO | float |
Histone_ChIP-seq_ENCSR000DVO |
Histone_ChIP-seq ENCSR000DVO [biosample_summary="Homo sapiens endothelial cell of umbilical vein newborn" and target="H3K27me3"]
|
 |
[1.82, 18.1] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF919ZCR.bed.gz | 1.82 MB | 297dcd77c7c84bb928ee130dd25345fd |
| ENCFF919ZCR.bed.gz.dvc | 101.0 B | a8c37e241b6783080c565e4a08443ae3 |
| ENCFF919ZCR.tabix.bed.gz | 958.69 KB | 411ea5ca7bcdd60ec8ad8c4176607ae7 |
| ENCFF919ZCR.tabix.bed.gz.dvc | 106.0 B | 5837f92bea16012acf7a7d13e6740850 |
| ENCFF919ZCR.tabix.bed.gz.tbi | 227.86 KB | a4bd484a9368daef961e3de7f7703cb9 |
| ENCFF919ZCR.tabix.bed.gz.tbi.dvc | 110.0 B | cf482eafec8668b9f35e90744baa982e |
| genomic_resource.yaml | 2.91 KB | 3d8e759f5557bc9af3d9785aad5ba8fb |
| genomic_resource_original.yaml | 2.77 KB | 2cbcf0a7e1d415ca604b65e84a64d6d7 |
| statistics/ |