Histone_ChIP-seq_ENCSR000DUA
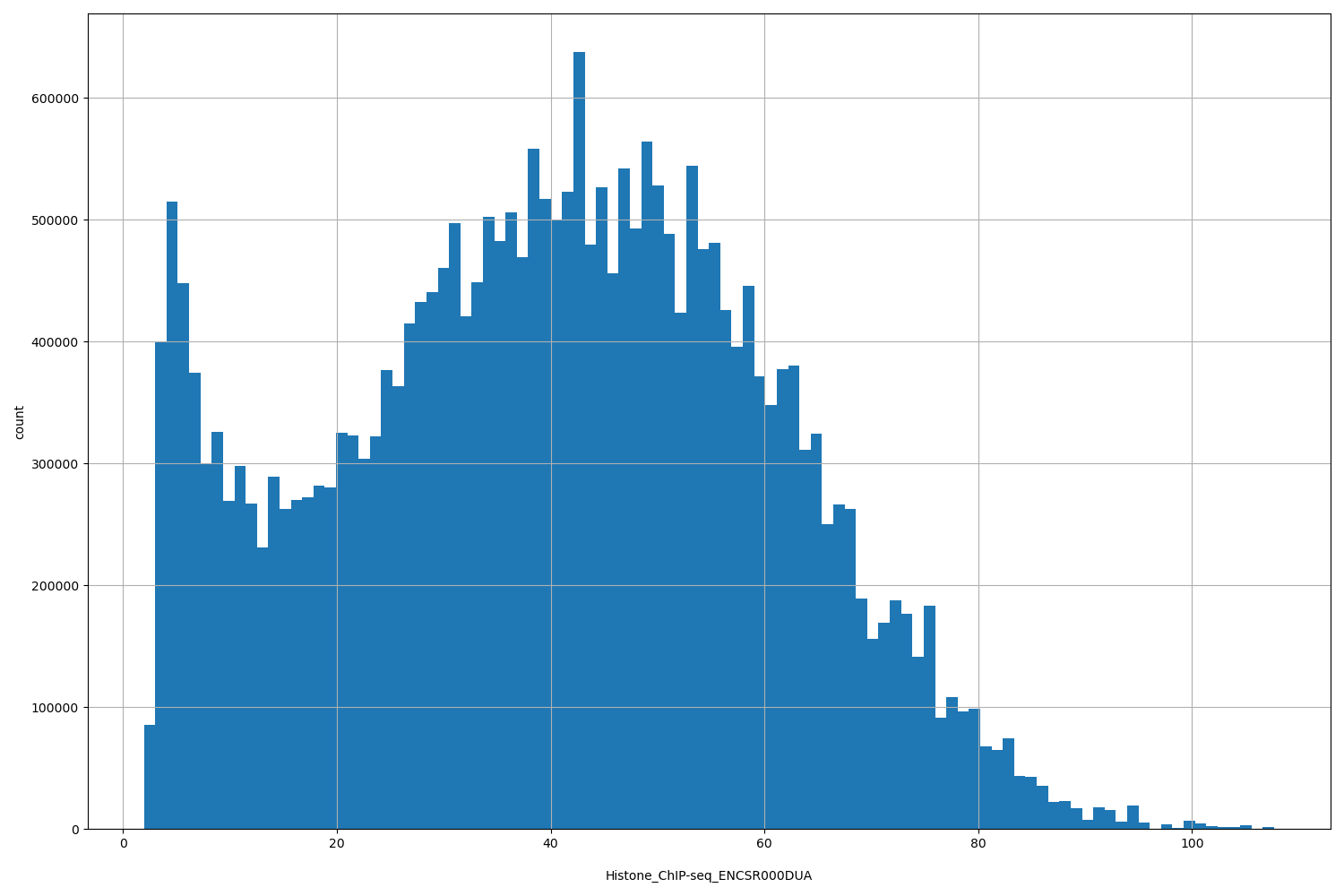
| Id: | Histone_ChIP-seq/ENCSR000DUA |
| Type: | position_score |
| Version: | 0 |
| Summary: |
HistoneChIP-seq ENCSR000DUA [biosamplesummary="Homo sapiens HeLa-S3 G1b phase" and target="H3K4me3"] |
| Description: |
status: released biological_replicates: Rep 1, Rep 2 summary: G1b phase output_type: replicated peaks audit_internal_action: Released analysis {ENCAN814YZN|/analyses/ENCAN814YZN/} has in progress subobject document {a6216f23-9635-413b-b9d0-0b35739abb65|/documents/a6216f23-9635-413b-b9d0-0b35739abb65/} audit_internal_action: Released analysis {ENCAN814YZN|/analyses/ENCAN814YZN/} has in progress subobject quality standard {encode4-histone-chip|/quality-standards/encode4-histone-chip/} audit_warning: Processed alignments file {ENCFF470WRZ|/files/ENCFF470WRZ/} processed by ChIP-seq ENCODE4 v1.5.1 GRCh38 pipeline has 13216300 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting H3K4me3-human and investigated as a narrow histone mark is 10 million usable fragments. The recommended value is > 20 million, but > 10 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/}) audit_warning: Processed alignments file {ENCFF847DCI|/files/ENCFF847DCI/} processed by ChIP-seq ENCODE4 v1.5.1 GRCh38 pipeline has 10771722 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting H3K4me3-human and investigated as a narrow histone mark is 10 million usable fragments. The recommended value is > 20 million, but > 10 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/}) |
| Labels: |
|
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| Histone_ChIP-seq_ENCSR000DUA | float |
Histone_ChIP-seq_ENCSR000DUA |
Histone_ChIP-seq ENCSR000DUA [biosample_summary="Homo sapiens HeLa-S3 G1b phase" and target="H3K4me3"]
|
 |
[1.94, 108] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF469OXD.bed.gz | 573.23 KB | 44c07bc1e0e662a6cb842849a20a44de |
| ENCFF469OXD.bed.gz.dvc | 100.0 B | 9324a574f3dab52800f4fe0013989e6b |
| ENCFF469OXD.tabix.bed.gz | 251.35 KB | b1ab0fb58bc878f08a81b01edc1f458b |
| ENCFF469OXD.tabix.bed.gz.dvc | 106.0 B | adb3fe88aa7132c6d38a089dc5718fc1 |
| ENCFF469OXD.tabix.bed.gz.tbi | 121.81 KB | 19525528df42d933561fa513735ee55b |
| ENCFF469OXD.tabix.bed.gz.tbi.dvc | 110.0 B | cb272de4b847f5aa17480c214e38e5c9 |
| genomic_resource.yaml | 2.89 KB | e5f9f0a46b588260f80ecf9db8475623 |
| genomic_resource_original.yaml | 2.77 KB | 0dff93b033d0aca71161f5914b1ab702 |
| statistics/ |