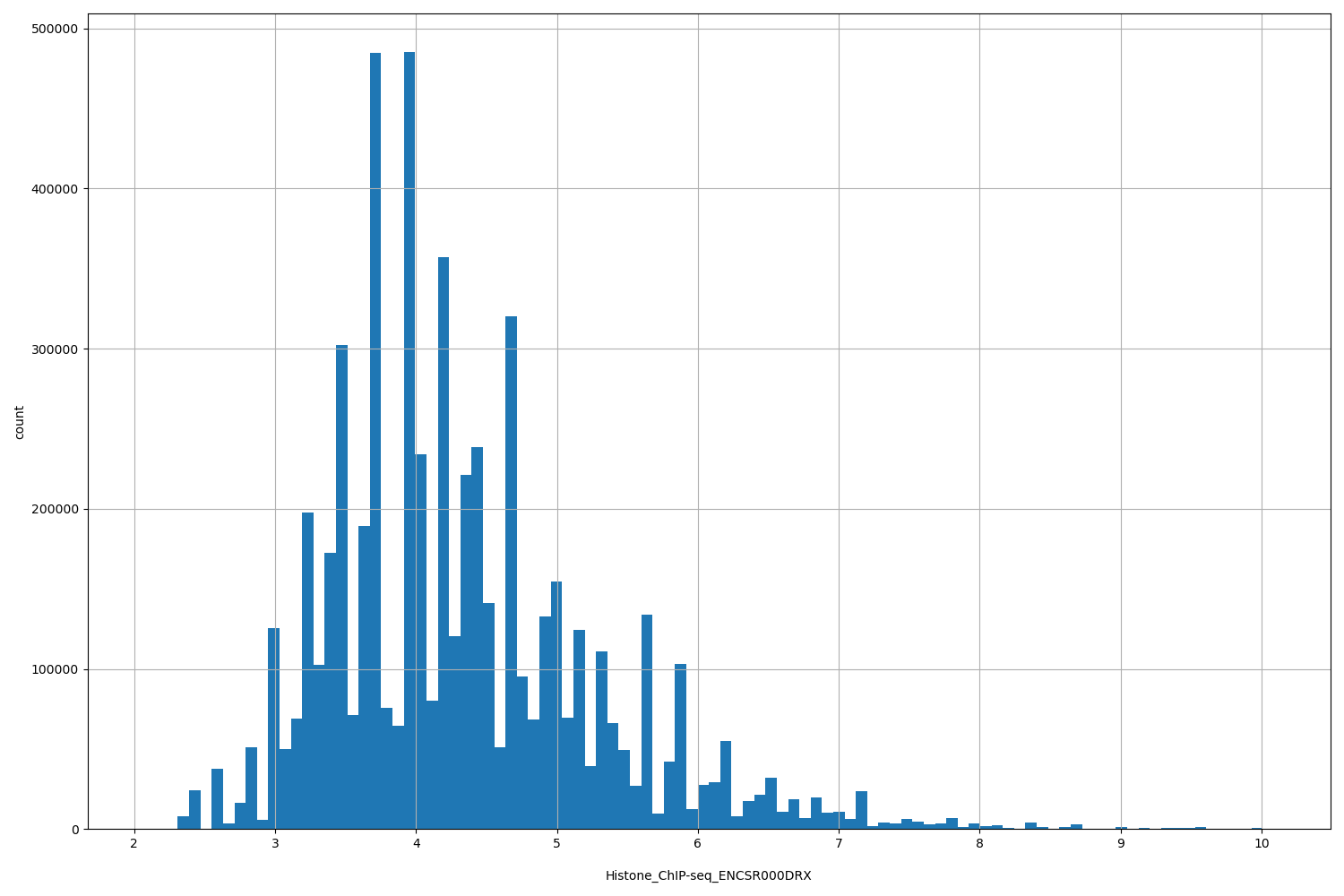
Histone_ChIP-seq_ENCSR000DRX
| Id: | Histone_ChIP-seq/ENCSR000DRX |
| Type: | position_score |
| Version: | 0 |
| Summary: |
HistoneChIP-seq ENCSR000DRX [biosamplesummary="Homo sapiens GM12878" and target="H3K27me3"] |
| Description: |
status: released biological_replicates: Rep 1, Rep 2 summary: output_type: replicated peaks audit_internal_action: Released analysis {ENCAN452ZJA|/analyses/ENCAN452ZJA/} has in progress subobject quality standard {encode4-histone-chip|/quality-standards/encode4-histone-chip/} audit_internal_action: Released analysis {ENCAN452ZJA|/analyses/ENCAN452ZJA/} has in progress subobject document {f674f66a-5fba-417c-9e20-d529a058193c|/documents/f674f66a-5fba-417c-9e20-d529a058193c/} audit_not_compliant: Processed alignments file {ENCFF265UBT|/files/ENCFF265UBT/} processed by ChIP-seq ENCODE4 v1.6.0 GRCh38 pipeline has 10821814 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting H3K27me3-human and investigated as a broad histone mark is 20 million usable fragments. The recommended value is > 45 million, but > 35 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/}) audit_not_compliant: Processed alignments file {ENCFF824VSE|/files/ENCFF824VSE/} processed by ChIP-seq ENCODE4 v1.6.0 GRCh38 pipeline has 14765423 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting H3K27me3-human and investigated as a broad histone mark is 20 million usable fragments. The recommended value is > 45 million, but > 35 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/}) |
| Labels: |
|
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| Histone_ChIP-seq_ENCSR000DRX | float |
Histone_ChIP-seq_ENCSR000DRX |
Histone_ChIP-seq ENCSR000DRX [biosample_summary="Homo sapiens GM12878" and target="H3K27me3"]
|
 |
[2.07, 10.1] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF695ETB.bed.gz | 435.34 KB | 850862bc88b52830686d31105976ff65 |
| ENCFF695ETB.bed.gz.dvc | 100.0 B | 3944ea0b03d87b5744719327f07d7d83 |
| ENCFF695ETB.tabix.bed.gz | 242.33 KB | d1e20a03914e965740f1c46f80acd9cd |
| ENCFF695ETB.tabix.bed.gz.dvc | 106.0 B | 83b8e77447b68eb555ffb39a6dc0c478 |
| ENCFF695ETB.tabix.bed.gz.tbi | 114.55 KB | ffb28061a5035dca2da6091c3ab72085 |
| ENCFF695ETB.tabix.bed.gz.tbi.dvc | 110.0 B | 1edfa47ab4b12c3ca5ae79f5dd9a5096 |
| genomic_resource.yaml | 2.91 KB | 370857c8954094f44a9fe87b639d6c6a |
| genomic_resource_original.yaml | 2.8 KB | 5bff29083f54a4ad404a71968a1fdcce |
| statistics/ |