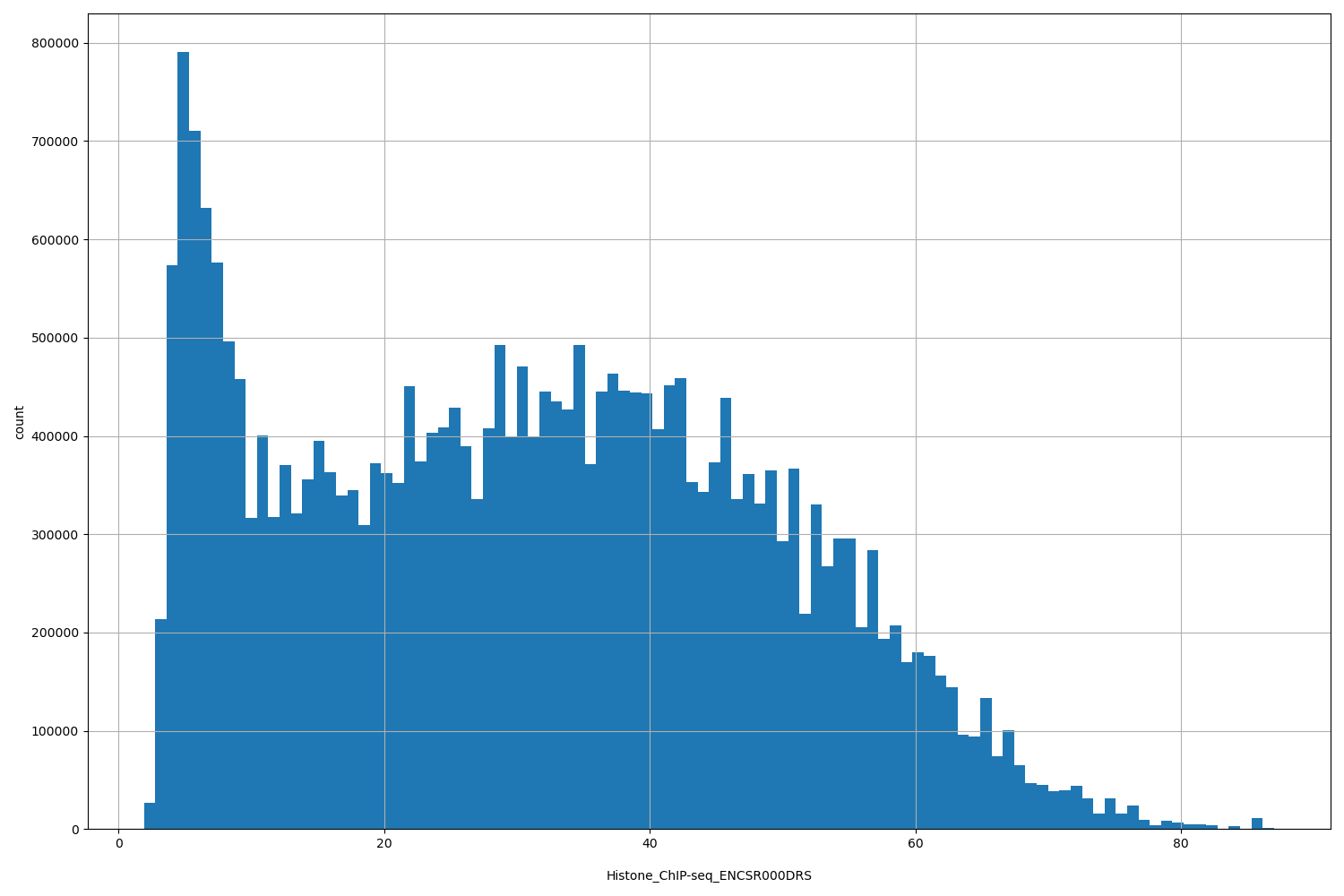
Histone_ChIP-seq_ENCSR000DRS
| Id: | Histone_ChIP-seq/ENCSR000DRS |
| Type: | position_score |
| Version: | 0 |
| Summary: |
HistoneChIP-seq ENCSR000DRS [biosamplesummary="Homo sapiens GM12875" and target="H3K4me3"] |
| Description: |
status: released biological_replicates: Rep 1 summary: output_type: pseudoreplicated peaks audit_internal_action: Archived analysis {ENCAN764WQM|/analyses/ENCAN764WQM/} has in progress subobject quality standard {encode3-histone-chip|/quality-standards/encode3-histone-chip/} audit_warning: Processed alignments file {ENCFF693JCT|/files/ENCFF693JCT/} processed by ChIP-seq ENCODE3 hg19 pipeline has 13008371 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting H3K4me3-human and investigated as a narrow histone mark is 10 million usable fragments. The recommended value is > 20 million, but > 10 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/}) |
| Labels: |
|
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| Histone_ChIP-seq_ENCSR000DRS | float |
Histone_ChIP-seq_ENCSR000DRS |
Histone_ChIP-seq ENCSR000DRS [biosample_summary="Homo sapiens GM12875" and target="H3K4me3"]
|
 |
[1.9, 87] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF770THI.bed.gz | 674.92 KB | e4b5582277d88b55bc18dc00d9526c1e |
| ENCFF770THI.bed.gz.dvc | 100.0 B | de34bfad93a17c2ca73226545bd8f7cf |
| ENCFF770THI.tabix.bed.gz | 297.95 KB | 475adaf40cc4374c666e43a34facb12b |
| ENCFF770THI.tabix.bed.gz.dvc | 106.0 B | 5796937d8c82486d1497c699d3de84bc |
| ENCFF770THI.tabix.bed.gz.tbi | 134.3 KB | db002392e46e67834f9a160859cf9d74 |
| ENCFF770THI.tabix.bed.gz.tbi.dvc | 110.0 B | a78014dd923ee4f707374b3d9bff90aa |
| genomic_resource.yaml | 2.07 KB | e5299fad443700fcf1e50eb4dad3aa75 |
| genomic_resource_original.yaml | 1.96 KB | aa0ce5241c8eabc1d0073ff23f96fec3 |
| statistics/ |