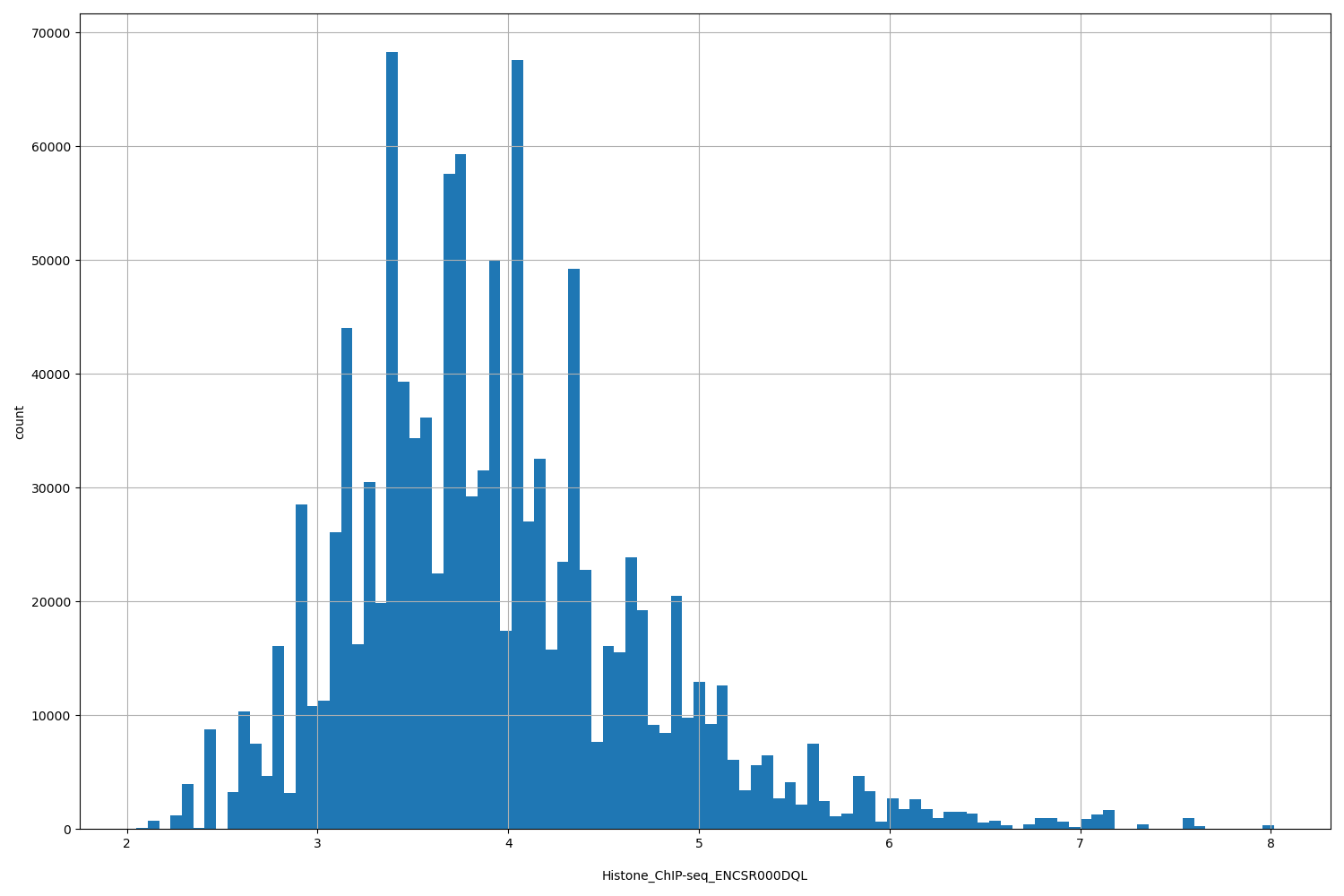
Histone_ChIP-seq_ENCSR000DQL
| Id: | Histone_ChIP-seq/ENCSR000DQL |
| Type: | position_score |
| Version: | 0 |
| Summary: |
HistoneChIP-seq ENCSR000DQL [biosamplesummary="Homo sapiens Caco-2" and target="H3K27me3"] |
| Description: |
status: released biological_replicates: Rep 1, Rep 2 summary: output_type: replicated peaks audit_internal_action: Released analysis {ENCAN923SXV|/analyses/ENCAN923SXV/} has in progress subobject quality standard {encode4-histone-chip|/quality-standards/encode4-histone-chip/} audit_internal_action: Released analysis {ENCAN923SXV|/analyses/ENCAN923SXV/} has in progress subobject document {06b95de9-03af-4bf2-9f73-c7b914c479a7|/documents/06b95de9-03af-4bf2-9f73-c7b914c479a7/} audit_not_compliant: Processed alignments file {ENCFF064IQV|/files/ENCFF064IQV/} processed by ChIP-seq ENCODE4 v1.6.0 GRCh38 pipeline has 14606302 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting H3K27me3-human and investigated as a broad histone mark is 20 million usable fragments. The recommended value is > 45 million, but > 35 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/}) audit_not_compliant: Processed alignments file {ENCFF957OES|/files/ENCFF957OES/} processed by ChIP-seq ENCODE4 v1.6.0 GRCh38 pipeline has 12760707 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting H3K27me3-human and investigated as a broad histone mark is 20 million usable fragments. The recommended value is > 45 million, but > 35 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/}) |
| Labels: |
|
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| Histone_ChIP-seq_ENCSR000DQL | float |
Histone_ChIP-seq_ENCSR000DQL |
Histone_ChIP-seq ENCSR000DQL [biosample_summary="Homo sapiens Caco-2" and target="H3K27me3"]
|
 |
[2.05, 8.01] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF649WIT.bed.gz | 115.67 KB | 510b9f844b0e7c9afb813c57a866d125 |
| ENCFF649WIT.bed.gz.dvc | 100.0 B | 7441a66ff6240ba4269b1db6a9aecbcd |
| ENCFF649WIT.tabix.bed.gz | 63.49 KB | f595d1507abbf1c1b7cb566c78f3e77e |
| ENCFF649WIT.tabix.bed.gz.dvc | 105.0 B | 3d079271c68527607fda0a1d5cef9cad |
| ENCFF649WIT.tabix.bed.gz.tbi | 45.8 KB | 6234b97a25152580ef1bdebc1d7e17e6 |
| ENCFF649WIT.tabix.bed.gz.tbi.dvc | 109.0 B | 653f18334d1c1b3537032a48d613917e |
| genomic_resource.yaml | 2.91 KB | b3d2d397c65513c189335381f3a1e681 |
| genomic_resource_original.yaml | 2.8 KB | 13053459b22c019aba28eceebe438b0a |
| statistics/ |