Histone_ChIP-seq_ENCSR000ATG
| Id: | Histone_ChIP-seq/ENCSR000ATG |
| Type: | position_score |
| Version: | 0 |
| Summary: |
HistoneChIP-seq ENCSR000ATG [biosamplesummary="Homo sapiens fibroblast of dermis NONE and female adult" and target="H2AFZ"] |
| Description: |
status: released biological_replicates: Rep 1, Rep 2 summary: female adult output_type: replicated peaks audit_internal_action: Archived analysis {ENCAN745IVJ|/analyses/ENCAN745IVJ/} has in progress subobject quality standard {encode3-histone-chip|/quality-standards/encode3-histone-chip/} audit_warning: Processed alignments file {ENCFF515NLW|/files/ENCFF515NLW/} processed by ChIP-seq ENCODE3 GRCh38 pipeline has 16857187 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting H2AFZ-human and investigated as a narrow histone mark is 10 million usable fragments. The recommended value is > 20 million, but > 10 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/}) audit_warning: Processed alignments file {ENCFF065NBX|/files/ENCFF065NBX/} processed by ChIP-seq ENCODE3 GRCh38 pipeline has 18063926 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting H2AFZ-human and investigated as a narrow histone mark is 10 million usable fragments. The recommended value is > 20 million, but > 10 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/}) audit_warning: PBC1 (PCR Bottlenecking Coefficient 1, M1/M_distinct) is the ratio of the number of genomic locations where exactly one read maps uniquely (M1) to the number of genomic locations where some reads map (M_distinct). A PBC1 value in the range 0 - 0.5 is severe bottlenecking, 0.5 - 0.8 is moderate bottlenecking, 0.8 - 0.9 is mild bottlenecking, and > 0.9 is no bottlenecking. PBC1 value > 0.9 is recommended, but > 0.8 is acceptable. ENCODE processed alignments file {ENCFF515NLW|/files/ENCFF515NLW/} processed by ChIP-seq ENCODE3 GRCh38 pipeline was generated from a library with PBC1 value of 0.88. audit_warning: PBC2 (PCR Bottlenecking Coefficient 2, M1/M2) is the ratio of the number of genomic locations where exactly one read maps uniquely (M1) to the number of genomic locations where two reads map uniquely (M2). A PBC2 value in the range 0 - 1 is severe bottlenecking, 1 - 3 is moderate bottlenecking, 3 - 10 is mild bottlenecking, > 10 is no bottlenecking. PBC2 value > 10 is recommended, but > 3 is acceptable. ENCODE processed alignments file {ENCFF515NLW|/files/ENCFF515NLW/} processed by ChIP-seq ENCODE3 GRCh38 pipeline was generated from a library with PBC2 value of 8.36. audit_warning: NRF (Non Redundant Fraction) is equal to the result of the division of the number of reads after duplicates removal by the total number of reads. An NRF value in the range 0 - 0.5 is poor complexity, 0.5 - 0.8 is moderate complexity, and > 0.8 high complexity. NRF value > 0.8 is recommended, but > 0.5 is acceptable. ENCODE processed alignments file {ENCFF065NBX|/files/ENCFF065NBX/} processed by ChIP-seq ENCODE3 GRCh38 pipeline was generated from a library with NRF value of 0.62. audit_warning: PBC1 (PCR Bottlenecking Coefficient 1, M1/M_distinct) is the ratio of the number of genomic locations where exactly one read maps uniquely (M1) to the number of genomic locations where some reads map (M_distinct). A PBC1 value in the range 0 - 0.5 is severe bottlenecking, 0.5 - 0.8 is moderate bottlenecking, 0.8 - 0.9 is mild bottlenecking, and > 0.9 is no bottlenecking. PBC1 value > 0.9 is recommended, but > 0.8 is acceptable. ENCODE processed alignments file {ENCFF065NBX|/files/ENCFF065NBX/} processed by ChIP-seq ENCODE3 GRCh38 pipeline was generated from a library with PBC1 value of 0.60. audit_warning: PBC2 (PCR Bottlenecking Coefficient 2, M1/M2) is the ratio of the number of genomic locations where exactly one read maps uniquely (M1) to the number of genomic locations where two reads map uniquely (M2). A PBC2 value in the range 0 - 1 is severe bottlenecking, 1 - 3 is moderate bottlenecking, 3 - 10 is mild bottlenecking, > 10 is no bottlenecking. PBC2 value > 10 is recommended, but > 3 is acceptable. ENCODE processed alignments file {ENCFF065NBX|/files/ENCFF065NBX/} processed by ChIP-seq ENCODE3 GRCh38 pipeline was generated from a library with PBC2 value of 2.35. |
| Labels: |
|
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| Histone_ChIP-seq_ENCSR000ATG | float |
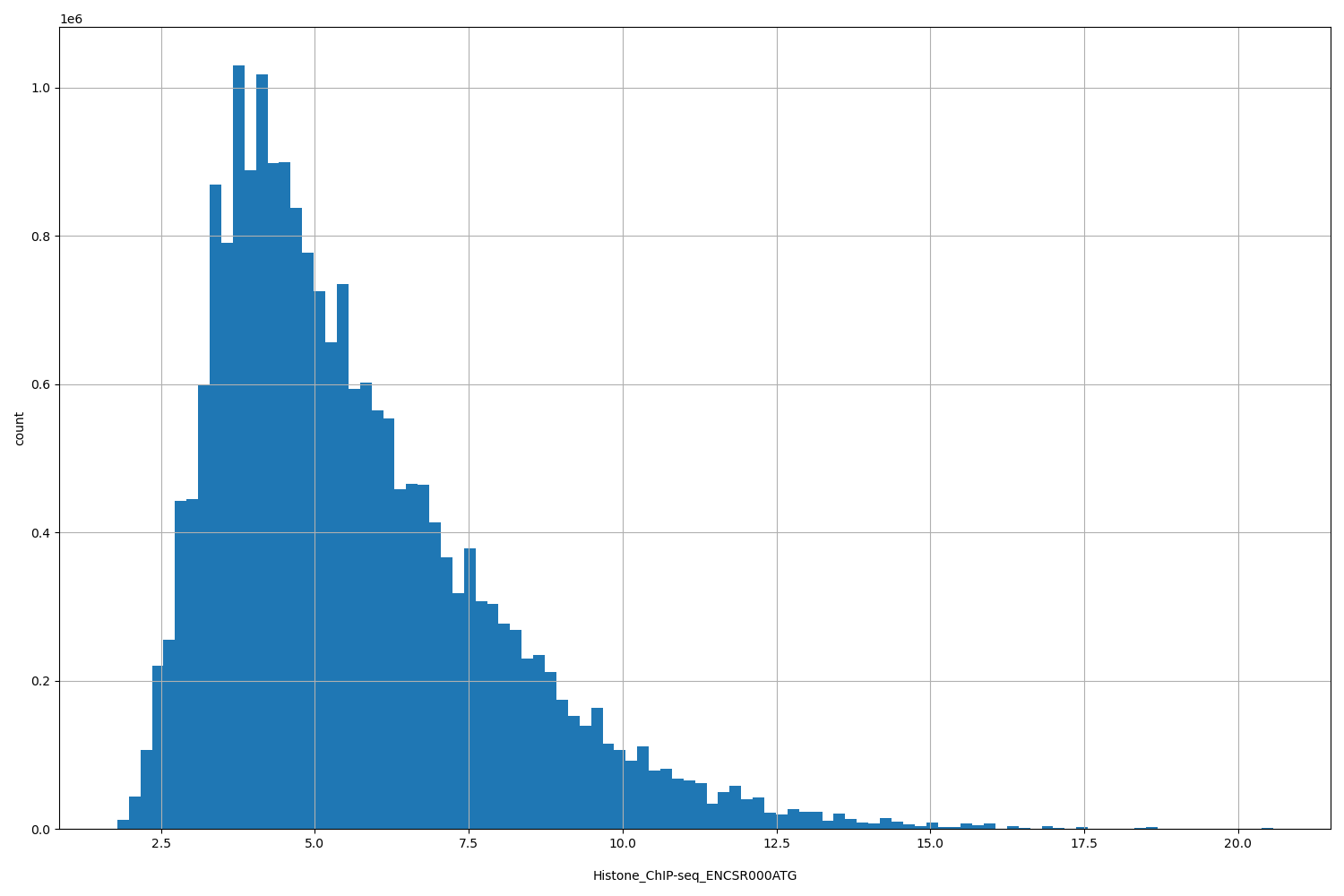
Histone_ChIP-seq_ENCSR000ATG |
Histone_ChIP-seq ENCSR000ATG [biosample_summary="Homo sapiens fibroblast of dermis NONE and female adult" and target="H2AFZ"]
|
 |
[1.79, 20.6] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF005LQD.bed.gz | 915.29 KB | 45496c7e31d95b85329970bc6b96dd06 |
| ENCFF005LQD.bed.gz.dvc | 100.0 B | c651762e5fdc4c6bbc8bb13a4ff5bd68 |
| ENCFF005LQD.tabix.bed.gz | 444.23 KB | 2a1d842449ce94bb16ffb19a027af9e2 |
| ENCFF005LQD.tabix.bed.gz.dvc | 106.0 B | 8f14db599b57035087cec4996e316e92 |
| ENCFF005LQD.tabix.bed.gz.tbi | 204.62 KB | 4d820cd75c0cf5281fe013e183c7effb |
| ENCFF005LQD.tabix.bed.gz.tbi.dvc | 110.0 B | 015ce208a469e3da81c4de90b2b785d0 |
| genomic_resource.yaml | 5.55 KB | d565515e7f8fdd4400ea5642af32b218 |
| genomic_resource_original.yaml | 5.42 KB | af0b84ff350b5a3404c381a046f8e33a |
| statistics/ |